Drug Information
Drug (ID: DG00063) and It's Reported Resistant Information
| Name |
Dalfopristin
|
||||
|---|---|---|---|---|---|
| Synonyms |
5-(2-DIETHYLAMINO-ETHANESULFONYL)-21-HYDROXY-10-ISOPROPYL-11,19-DIMETHYL-9,26-DIOXA-3,15,28-TRIAZA-TRICYCLO[23.2.1.00,255]OCTACOSA-1(27),12,17,19,25(28)-PENTAENE-2,8,14,23-TETRAONE
Click to Show/Hide
|
||||
| Indication |
In total 1 Indication(s)
|
||||
| Structure |
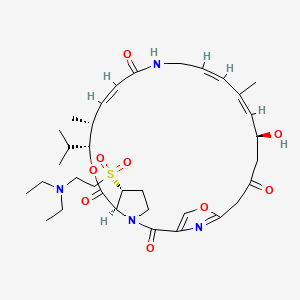
|
||||
| Drug Resistance Disease(s) |
Disease(s) with Clinically Reported Resistance for This Drug
(9 diseases)
[3]
[3]
[3]
[3]
[3]
[3]
[3]
[3]
|
||||
| Target | Bacterial 50S ribosomal RNA (Bact 50S rRNA) | NOUNIPROTAC | [1] | ||
| Click to Show/Hide the Molecular Information and External Link(s) of This Drug | |||||
| Formula |
C34H50N4O9S
|
||||
| IsoSMILES |
CCN(CC)CCS(=O)(=O)[C@@H]1CCN2[C@H]1C(=O)O[C@@H]([C@@H](/C=C/C(=O)NC/C=C/C(=C/[C@H](CC(=O)CC3=NC(=CO3)C2=O)O)/C)C)C(C)C
|
||||
| InChI |
1S/C34H50N4O9S/c1-7-37(8-2)16-17-48(44,45)28-13-15-38-31(28)34(43)47-32(22(3)4)24(6)11-12-29(41)35-14-9-10-23(5)18-25(39)19-26(40)20-30-36-27(21-46-30)33(38)42/h9-12,18,21-22,24-25,28,31-32,39H,7-8,13-17,19-20H2,1-6H3,(H,35,41)/b10-9+,12-11+,23-18+/t24-,25-,28-,31-,32-/m1/s1
|
||||
| InChIKey |
SUYRLXYYZQTJHF-VMBLUXKRSA-N
|
||||
| PubChem CID | |||||
| ChEBI ID | |||||
| TTD Drug ID | |||||
| DrugBank ID | |||||
Type(s) of Resistant Mechanism of This Drug
Drug Resistance Data Categorized by Their Corresponding Diseases
ICD-01: Infectious/parasitic diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: ABC transporter ATP-binding protein (ABCP) | [1], [2] | |||
| Resistant Disease | Bacterial infection [ICD-11: 1A00-1C4Z] | |||
| Molecule Alteration | Missense mutation | p.T450I |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli TOP10 | 83333 | ||
| Enterococcus faecium HM1070 | 1352 | |||
| Enterococcus faecium UCN80 | 1352 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay | |||
| Mechanism Description | ABC systems constitute one of the largest families of proteins, with most of them being involved in import and export, often called ABC transporters.Several of these class 2 ABC systems have been involved in MLS resistance, such as Msr-, Vga-, or Lsa-like proteins.The observed profile of cross-resistance to lincosamides, streptogramins A, and pleuromutilins conferred by Eat(A)v was similar to those conferred by other Lsa-like proteins. | |||
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: ABC protein lsaC (lsaC-Unclear) | [3] | |||
| Resistant Disease | Streptococcus agalactiae infection [ICD-11: 1B21.2] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli TOP10 | 83333 | ||
| Staphylococcus aureus ATCC 29213 | 1280 | |||
| Streptococcus agalactiae UCN70 | 1311 | |||
| Streptococcus agalactiae isolates | 1311 | |||
| Streptococcus agalactiae BM132 | 1319 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay; Allelic frequency measurement assay | |||
| Experiment for Drug Resistance |
Broth microdilution method assay | |||
| Mechanism Description | Expression of this novel gene, named lsa(C), in S. agalactiae BM132 after cloning led to an increase in MICs of lincomycin (0.06 to 4 ug/ml), clindamycin (0.03 to 2 ug/ml), dalfopristin (2 to >32 ug/ml), and tiamulin (0.12 to 32 ug/ml), whereas no change in MICs of erythromycin (0.06 ug/ml), azithromycin (0.03 ug/ml), spiramycin (0.25 ug/ml), telithromycin (0.03 ug/ml), and quinupristin (8 ug/ml) was observed. The phenotype was renamed the LS(A)P phenotype on the basis of cross-resistance to lincosamides, streptogramins A, and pleuromutilins. | |||
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: ABC protein lsaC (lsaC-Unclear) | [3] | |||
| Resistant Disease | Streptococcus agalactiae infection [ICD-11: 1B21.2] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli TOP10 | 83333 | ||
| Staphylococcus aureus ATCC 29213 | 1280 | |||
| Streptococcus agalactiae UCN70 | 1311 | |||
| Streptococcus agalactiae isolates | 1311 | |||
| Streptococcus agalactiae BM132 | 1319 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay; Allelic frequency measurement assay | |||
| Experiment for Drug Resistance |
Broth microdilution method assay | |||
| Mechanism Description | Expression of this novel gene, named lsa(C), in S. agalactiae BM132 after cloning led to an increase in MICs of lincomycin (0.06 to 4 ug/ml), clindamycin (0.03 to 2 ug/ml), dalfopristin (2 to >32 ug/ml), and tiamulin (0.12 to 32 ug/ml), whereas no change in MICs of erythromycin (0.06 ug/ml), azithromycin (0.03 ug/ml), spiramycin (0.25 ug/ml), telithromycin (0.03 ug/ml), and quinupristin (8 ug/ml) was observed. The phenotype was renamed the LS(A)P phenotype on the basis of cross-resistance to lincosamides, streptogramins A, and pleuromutilins. | |||
ICD-11: Circulatory system diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: ABC protein lsaC (lsaC-Unclear) | [3] | |||
| Resistant Disease | Streptococcus agalactiae infection [ICD-11: 1B21.2] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli TOP10 | 83333 | ||
| Staphylococcus aureus ATCC 29213 | 1280 | |||
| Streptococcus agalactiae UCN70 | 1311 | |||
| Streptococcus agalactiae isolates | 1311 | |||
| Streptococcus agalactiae BM132 | 1319 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay; Allelic frequency measurement assay | |||
| Experiment for Drug Resistance |
Broth microdilution method assay | |||
| Mechanism Description | Expression of this novel gene, named lsa(C), in S. agalactiae BM132 after cloning led to an increase in MICs of lincomycin (0.06 to 4 ug/ml), clindamycin (0.03 to 2 ug/ml), dalfopristin (2 to >32 ug/ml), and tiamulin (0.12 to 32 ug/ml), whereas no change in MICs of erythromycin (0.06 ug/ml), azithromycin (0.03 ug/ml), spiramycin (0.25 ug/ml), telithromycin (0.03 ug/ml), and quinupristin (8 ug/ml) was observed. The phenotype was renamed the LS(A)P phenotype on the basis of cross-resistance to lincosamides, streptogramins A, and pleuromutilins. | |||
ICD-12: Respiratory system diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: ABC protein lsaC (lsaC-Unclear) | [3] | |||
| Resistant Disease | Streptococcus agalactiae infection [ICD-11: 1B21.2] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli TOP10 | 83333 | ||
| Staphylococcus aureus ATCC 29213 | 1280 | |||
| Streptococcus agalactiae UCN70 | 1311 | |||
| Streptococcus agalactiae isolates | 1311 | |||
| Streptococcus agalactiae BM132 | 1319 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay; Allelic frequency measurement assay | |||
| Experiment for Drug Resistance |
Broth microdilution method assay | |||
| Mechanism Description | Expression of this novel gene, named lsa(C), in S. agalactiae BM132 after cloning led to an increase in MICs of lincomycin (0.06 to 4 ug/ml), clindamycin (0.03 to 2 ug/ml), dalfopristin (2 to >32 ug/ml), and tiamulin (0.12 to 32 ug/ml), whereas no change in MICs of erythromycin (0.06 ug/ml), azithromycin (0.03 ug/ml), spiramycin (0.25 ug/ml), telithromycin (0.03 ug/ml), and quinupristin (8 ug/ml) was observed. The phenotype was renamed the LS(A)P phenotype on the basis of cross-resistance to lincosamides, streptogramins A, and pleuromutilins. | |||
ICD-15: Musculoskeletal/connective-tissue diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: ABC protein lsaC (lsaC-Unclear) | [3] | |||
| Resistant Disease | Streptococcus agalactiae infection [ICD-11: 1B21.2] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli TOP10 | 83333 | ||
| Staphylococcus aureus ATCC 29213 | 1280 | |||
| Streptococcus agalactiae UCN70 | 1311 | |||
| Streptococcus agalactiae isolates | 1311 | |||
| Streptococcus agalactiae BM132 | 1319 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay; Allelic frequency measurement assay | |||
| Experiment for Drug Resistance |
Broth microdilution method assay | |||
| Mechanism Description | Expression of this novel gene, named lsa(C), in S. agalactiae BM132 after cloning led to an increase in MICs of lincomycin (0.06 to 4 ug/ml), clindamycin (0.03 to 2 ug/ml), dalfopristin (2 to >32 ug/ml), and tiamulin (0.12 to 32 ug/ml), whereas no change in MICs of erythromycin (0.06 ug/ml), azithromycin (0.03 ug/ml), spiramycin (0.25 ug/ml), telithromycin (0.03 ug/ml), and quinupristin (8 ug/ml) was observed. The phenotype was renamed the LS(A)P phenotype on the basis of cross-resistance to lincosamides, streptogramins A, and pleuromutilins. | |||
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: ABC protein lsaC (lsaC-Unclear) | [3] | |||
| Resistant Disease | Streptococcus agalactiae inection [ICD-11: FB84.0] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli TOP10 | 83333 | ||
| Staphylococcus aureus ATCC 29213 | 1280 | |||
| Streptococcus agalactiae UCN70 | 1311 | |||
| Streptococcus agalactiae isolates | 1311 | |||
| Streptococcus agalactiae BM132 | 1319 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay; Allelic frequency measurement assay | |||
| Experiment for Drug Resistance |
Broth microdilution method assay | |||
| Mechanism Description | Expression of this novel gene, named lsa(C), in S. agalactiae BM132 after cloning led to an increase in MICs of lincomycin (0.06 to 4 ug/ml), clindamycin (0.03 to 2 ug/ml), dalfopristin (2 to >32 ug/ml), and tiamulin (0.12 to 32 ug/ml), whereas no change in MICs of erythromycin (0.06 ug/ml), azithromycin (0.03 ug/ml), spiramycin (0.25 ug/ml), telithromycin (0.03 ug/ml), and quinupristin (8 ug/ml) was observed. The phenotype was renamed the LS(A)P phenotype on the basis of cross-resistance to lincosamides, streptogramins A, and pleuromutilins. | |||
ICD-16: Genitourinary system diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: ABC protein lsaC (lsaC-Unclear) | [3] | |||
| Resistant Disease | Klebsiella pneumoniae [ICD-11: CA40.0] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli TOP10 | 83333 | ||
| Staphylococcus aureus ATCC 29213 | 1280 | |||
| Streptococcus agalactiae UCN70 | 1311 | |||
| Streptococcus agalactiae isolates | 1311 | |||
| Streptococcus agalactiae BM132 | 1319 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay; Allelic frequency measurement assay | |||
| Experiment for Drug Resistance |
Broth microdilution method assay | |||
| Mechanism Description | Expression of this novel gene, named lsa(C), in S. agalactiae BM132 after cloning led to an increase in MICs of lincomycin (0.06 to 4 ug/ml), clindamycin (0.03 to 2 ug/ml), dalfopristin (2 to >32 ug/ml), and tiamulin (0.12 to 32 ug/ml), whereas no change in MICs of erythromycin (0.06 ug/ml), azithromycin (0.03 ug/ml), spiramycin (0.25 ug/ml), telithromycin (0.03 ug/ml), and quinupristin (8 ug/ml) was observed. The phenotype was renamed the LS(A)P phenotype on the basis of cross-resistance to lincosamides, streptogramins A, and pleuromutilins. | |||
ICD-21: Symptoms/clinical signs/unclassified clinical findings
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: ABC protein lsaC (lsaC-Unclear) | [3] | |||
| Resistant Disease | Streptococcus agalactiae infection [ICD-11: 1B21.2] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli TOP10 | 83333 | ||
| Staphylococcus aureus ATCC 29213 | 1280 | |||
| Streptococcus agalactiae UCN70 | 1311 | |||
| Streptococcus agalactiae isolates | 1311 | |||
| Streptococcus agalactiae BM132 | 1319 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay; Allelic frequency measurement assay | |||
| Experiment for Drug Resistance |
Broth microdilution method assay | |||
| Mechanism Description | Expression of this novel gene, named lsa(C), in S. agalactiae BM132 after cloning led to an increase in MICs of lincomycin (0.06 to 4 ug/ml), clindamycin (0.03 to 2 ug/ml), dalfopristin (2 to >32 ug/ml), and tiamulin (0.12 to 32 ug/ml), whereas no change in MICs of erythromycin (0.06 ug/ml), azithromycin (0.03 ug/ml), spiramycin (0.25 ug/ml), telithromycin (0.03 ug/ml), and quinupristin (8 ug/ml) was observed. The phenotype was renamed the LS(A)P phenotype on the basis of cross-resistance to lincosamides, streptogramins A, and pleuromutilins. | |||
References
If you find any error in data or bug in web service, please kindly report it to Dr. Sun and Dr. Yu.
