Drug Information
Drug (ID: DG00121) and It's Reported Resistant Information
| Name |
Ticarcillin
|
||||
|---|---|---|---|---|---|
| Synonyms |
TIPC; Ticarcilina; Ticarcilline; Ticarcillinum; Ticarcillin Supplement; Ticillin [veterinary]; BRL-2288; Ticarcilina[INN-Spanish]; Ticarcillin (INN); Ticarcillin [INN:BAN]; Ticarcilline [INN-French]; Ticarcillinum [INN-Latin]; Ticillin [veterinary] (TN); Timentin (TN); A-carboxy-3-thienylmethylpenicillin; Alpha-carboxy-3-thienylmethylpenicillin; (2S,5R,6R)-6-[[(2R)-3-hydroxy-3-oxo-2-thiophen-3-ylpropanoyl]amino]-3,3-dimethyl-7-oxo-4-thia-1-azabicyclo[3.2.0]heptane-2-carboxylic acid; (2S,5R,6R)-6-{[(2R)-2-carboxy-2-(3-thienyl)acetyl]amino}-3,3-dimethyl-7-oxo-4-thia-1-azabicyclo[3.2.0]heptane-2-carboxylic acid; (2S,5R,6R)-6-{[(2R)-2-carboxy-2-thiophen-3-ylacetyl]amino}-3,3-dimethyl-7-oxo-4-thia-1-azabicyclo[3.2.0]heptane-2-carboxylic acid; 6beta-{[(2R)-2-carboxy-2-thiophen-3-ylacetyl]amino}-2,2-dimethylpenam-3alpha-carboxylic acid
Click to Show/Hide
|
||||
| Indication |
In total 1 Indication(s)
|
||||
| Structure |
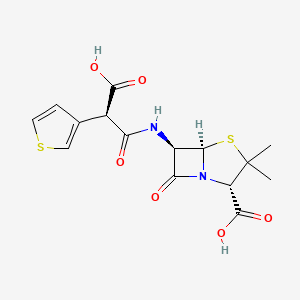
|
||||
| Drug Resistance Disease(s) |
Disease(s) with Clinically Reported Resistance for This Drug
(3 diseases)
[7]
|
||||
| Target | Bacterial Penicillin binding protein (Bact PBP) | NOUNIPROTAC | [1] | ||
| Click to Show/Hide the Molecular Information and External Link(s) of This Drug | |||||
| Formula |
C15H16N2O6S2
|
||||
| IsoSMILES |
CC1([C@@H](N2[C@H](S1)[C@@H](C2=O)NC(=O)[C@@H](C3=CSC=C3)C(=O)O)C(=O)O)C
|
||||
| InChI |
1S/C15H16N2O6S2/c1-15(2)9(14(22)23)17-11(19)8(12(17)25-15)16-10(18)7(13(20)21)6-3-4-24-5-6/h3-5,7-9,12H,1-2H3,(H,16,18)(H,20,21)(H,22,23)/t7-,8-,9+,12-/m1/s1
|
||||
| InChIKey |
OHKOGUYZJXTSFX-KZFFXBSXSA-N
|
||||
| PubChem CID | |||||
| ChEBI ID | |||||
| TTD Drug ID | |||||
| INTEDE ID | |||||
| DrugBank ID | |||||
Type(s) of Resistant Mechanism of This Drug
Drug Resistance Data Categorized by Their Corresponding Diseases
ICD-01: Infectious/parasitic diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Beta-lactamase (BLA) | [2], [3] | |||
| Resistant Disease | Anaerobic Bacterial infection [ICD-11: 1A00-1A09] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli HB101 | 634468 | ||
| Escherichia coli JM109 | 562 | |||
| Acidaminococcus fermentans RYC-MR95 | 905 | |||
| Acidaminococcus fermentans RYC4093 | 905 | |||
| Acidaminococcus fermentans RYC4356 | 905 | |||
| Escherichia coli RYC1000 | 562 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay | |||
| Experiment for Drug Resistance |
Agar dilution method assay; broth microdilution method assay | |||
| Mechanism Description | A. intestini is the first Gram-negative coccus with demonstrated resistance to beta-lactam antibiotics. The reference genome of the A. intestini strain RyC-MR95, which was isolated from a perianal abscess of a European male diabetic patient, contains the aci1 gene, which encodes the ACI-1 class A beta-lactamase that confers resistance to penicillins and extended-spectrum cephalosporins. | |||
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Metallo-beta-lactamase NDM-4 (NDM4) | [1] | |||
| Resistant Disease | Bacterial infection [ICD-11: 1A00-1C4Z] | |||
| Molecule Alteration | Missense mutation | p.M154L |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli I5 | 562 | ||
| Experiment for Molecule Alteration |
Whole genome sequence assay; Allelic frequency measurement assay | |||
| Experiment for Drug Resistance |
Disk diffusion test assay; E-strip test assay | |||
| Mechanism Description | A clinical Escherichia coli isolate resistant to all Beta-lactams, including carbapenems, expressed a novel metallo-Beta-lactamase (MBL), NDM-4, differing from NDM-1 by a single amino acid substitution (Met154Leu). NDM-4 possessed increased hydrolytic activity toward carbapenems and several cephalosporins compared to that of NDM-1. | |||
| Key Molecule: Beta-lactamase (BLA) | [8] | |||
| Resistant Disease | Bacterial infection [ICD-11: 1A00-1C4Z] | |||
| Molecule Alteration | Missense mutation | p.Y104A+p.N110D+p.E175Q+p.S179A |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli TOP10 | 83333 | ||
| Acinetobacter baumannii CIP70.10 | 470 | |||
| Klebsiella pneumoniae kP3 | 1290996 | |||
| Pseudomonas aeruginosa PU21 | 287 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay; Allelic frequency measurement assay | |||
| Experiment for Drug Resistance |
MIC assay | |||
| Mechanism Description | K. pneumoniae kP3 was resistant to all Beta-lactams, including carbapenems, and expressed the carbapenem-hydrolyzing Beta-lactamase OXA-181, which differs from OXA-48 by four amino acid substitutions. Compared to OXA-48, OXA-181 possessed a very similar hydrolytic profile. | |||
| Key Molecule: Beta-lactamase (BLA) | [6], [9] | |||
| Resistant Disease | Bacterial infection [ICD-11: 1A00-1C4Z] | |||
| Molecule Alteration | Missense mutation | p.D240G |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli | 668369 | ||
| Escherichia coli Gre-1 | 562 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay | |||
| Experiment for Drug Resistance |
Agar dilution method assay | |||
| Mechanism Description | The first extended-spectrum Beta-lactamase (ESBL) of the CTX-M type (MEN-1/CTX-M-1) was reported at the beginning of the 1990s.CTX-M-27 differed from CTX-M-14 only by the substitution D240G and was the third CTX-M enzyme harbouring this mutation after CTX-M-15 and CTX-M-16. The Gly-240-harbouring enzyme CTX-M-27 conferred to Escherichia coli higher MICs of ceftazidime (MIC, 8 versus 1 mg/L) than did the Asp-240-harbouring CTX-M-14 enzyme. | |||
| Key Molecule: Beta-lactamase (BLA) | [4], [5], [6] | |||
| Resistant Disease | Bacterial infection [ICD-11: 1A00-1C4Z] | |||
| Molecule Alteration | Missense mutation | p.D240G |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli DH10B | 316385 | ||
| Citrobacter freundii 2526/96 | 546 | |||
| Escherichia coli isolates | 562 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay | |||
| Experiment for Drug Resistance |
Agar dilution method assay | |||
| Mechanism Description | We have reported recently the DNA sequence of another Beta-lactamase, CTX- M-15, from Indian enterobacterial isolates that were resistant to both cefotaxime and ceftazidime.CTX-M-15 has a single amino acid change [Asp-240-Gly (Ambler numbering)]7 compared with CTX-M-3. | |||
ICD-12: Respiratory system diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Bcr/CflA family efflux transporter (BCML) | [7] | |||
| Resistant Disease | Klebsiella pneumoniae infection [ICD-11: CA40.1] | |||
| Molecule Alteration | Expression | Inherence |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli DH10B | 316385 | ||
| Escherichia coli strain NCTC 50192 | 562 | |||
| Klebsiella pneumoniae strain ORI-1 | 573 | |||
| Experiment for Molecule Alteration |
PCR and hybridization experiments assay | |||
| Experiment for Drug Resistance |
Agar dilution technique assay | |||
| Mechanism Description | Klebsiella pneumoniae ORI-1 strain harbored a ca. 140-kb nontransferable plasmid, pTk1, that conferred an extended-spectrum cephalosporin resistance profile antagonized by the addition of clavulanic acid, tazobactam, or imipenem. The gene for GES-1 (Guiana extended-spectrum beta-lactamase) was cloned, and its protein was expressed in Escherichia coli DH10B, where this pI-5. 8 beta-lactamase of a ca. 31-kDa molecular mass conferred resistance to oxyimino cephalosporins (mostly to ceftazidime). GES-1 is weakly related to the other plasmid-located Ambler class A extended-spectrum beta-lactamases (ESBLs). | |||
| Key Molecule: Bcr/CflA family efflux transporter (BCML) | [7] | |||
| Resistant Disease | Klebsiella pneumoniae infection [ICD-11: CA40.1] | |||
| Molecule Alteration | Expression | Acquired |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli DH10B | 316385 | ||
| Escherichia coli strain NCTC 50192 | 562 | |||
| Klebsiella pneumoniae strain ORI-1 | 573 | |||
| Experiment for Molecule Alteration |
PCR and hybridization experiments assay | |||
| Experiment for Drug Resistance |
Agar dilution technique assay | |||
| Mechanism Description | Beta-Lactam MICs for k. pneumoniae ORI-1 and Escherichia coli DH10B harboring either the natural plasmid pTk1 or the recombinant plasmid pC1 were somewhat similar and might indicate the presence of an ESBL. In all cases, the ceftazidime MICs were higher than those of cefotaxime and aztreonam. Beta-Lactam MICs were always lowered by the addition of clavulanic acid or tazobactam, less so by sulbactam, and uncommonly by imipenem. | |||
| Drug Sensitivity Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Bcr/CflA family efflux transporter (BCML) | [7] | |||
| Sensitive Disease | Klebsiella pneumoniae infection [ICD-11: CA40.1] | |||
| Molecule Alteration | Expression | Antagonism |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli DH10B | 316385 | ||
| Escherichia coli strain NCTC 50192 | 562 | |||
| Klebsiella pneumoniae strain ORI-1 | 573 | |||
| Experiment for Molecule Alteration |
PCR and hybridization experiments assay | |||
| Experiment for Drug Resistance |
Agar dilution technique assay | |||
| Mechanism Description | Inhibition studies, as measured by IC50 values with benzylpenicillin as the substrate, showed that GES-1 was inhibited by clavulanic acid (5 uM) and tazobactam (2.5 uM) and strongly inhibited by imipenem (0.1 uM). Beta-Lactam MICs were always lowered by the addition of clavulanic acid or tazobactam, less so by sulbactam, and uncommonly by imipenem. | |||
References
If you find any error in data or bug in web service, please kindly report it to Dr. Sun and Dr. Yu.
