Drug Information
Drug (ID: DG00136) and It's Reported Resistant Information
| Name |
Isoniazid
|
||||
|---|---|---|---|---|---|
| Synonyms |
Abdizide; Andrazide; Anidrasona; Antimicina; Antituberkulosum; Armacide; Armazid; Armazide; Atcotibine; Azuren; Bacillen; Bacillin; Cedin; Cemidon; Chemiazid; Chemidon; Continazine; Cortinazine; Cotinazin; Cotinizin; Defonin; Dibutin; Diforin; Dinacrin; Ditubin; Ebidene; Eralon; Ertuban; Eutizon; Evalon; Fetefu; Fimalene; HIA; Hidranizil; Hidrasonil; Hidrulta; Hidrun; Hycozid; Hydra; Hydrazid; Hydrazide; Hyozid; Hyzyd; INH; Idrazil; Inah; Inizid; Iscotin; Isidrina; Ismazide; Isobicina; Isocid; Isocidene; Isocotin; Isohydrazide; Isokin; Isolyn; Isonerit; Isonex; Isoniacid; Isoniazida; Isoniazide; Isoniazidum; Isonicazide; Isonicid; Isonico; Isonicotan; Isonicotil; Isonicotinhydrazid; Isonicotinohydrazide; Isonide; Isonidrin; Isonikazid; Isonilex; Isonin; Isonindon; Isonirit; Isoniton; Isonizida; Isonizide; Isotamine; Isotebe; Isotebezid; Isotinyl; Isozid; Isozide; Isozyd; LANIZID; Laniazid; Laniozid; Mybasan; Neoteben; Neoxin; Neumandin; Nevin; Niadrin; Nicazide; Nicetal; Nicizina; Niconyl; Nicotibina; Nicotibine; Nicotisan; Nicozide; Nidaton; Nidrazid; Nikozid; Niplen; Nitadon; Niteban; Nydrazid; Nyscozid; Pelazid; Percin; Phthisen; Pycazide; Pyreazid; Pyricidin; Pyridicin; Pyrizidin; Raumanon; Razide; Retozide; Rimicid; Rimifon; Rimiphone; Rimitsid; Robiselin; Robisellin; Roxifen; Sanohidrazina; Sauterazid; Sauterzid; Stanozide; Tebecid; Tebemid; Tebenic; Tebexin; Tebilon; Tebos; Teebaconin; Tekazin; Tibazide; Tibemid; Tibiazide; Tibinide; Tibison; Tibivis; Tibizide; Tibusan; Tisin; Tisiodrazida; Tizide; Tubazid; Tubazide; Tubeco; Tubecotubercid; Tubercid; Tuberian; Tubicon; Tubilysin; Tubizid; Tubomel; Tyvid; Unicocyde; Unicozyde; Vazadrine; Vederon; Zidafimia; Zinadon; Zonazide; Hid rasonil; Isoco tin; Isoniazid SA; Isozid e; Nidra zid; Rimif on; BP 5015; Bp 5 015; FSR 3; I0138; INHd20; L 1945; Nitebannsc 9659; Preparation 6424; RP 5015; AZT + Isoniazid; Cedin (Aerosol); Dow-Isoniazid; FRS-3; FSR-3; Ido-tebin; In-73; Inh-Burgthal; Isoniazid & EEP; Isoniazid & Propolis; Laniazid (TN); Neo-Tizide; Nydrazid (TN); RP-5015; TB-Phlogin; TB-Razide; TB-Vis; Usaf cb-2; I.A.I; RU-EF-Tb; RY-EF-Tb; I.A.I.
Click to Show/Hide
|
||||
| Indication |
In total 1 Indication(s)
|
||||
| Structure |
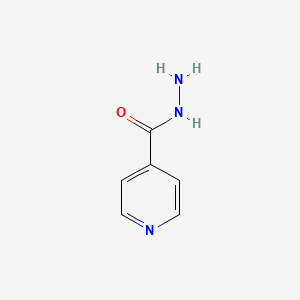
|
||||
| Drug Resistance Disease(s) |
Disease(s) with Resistance Information Discovered by Cell Line Test for This Drug
(3 diseases)
[2]
[3]
[6]
Disease(s) with Clinically Reported Resistance for This Drug
(4 diseases)
[4]
[5]
[7]
[8]
Disease(s) with Resistance Information Validated by in-vivo Model for This Drug
(2 diseases)
[9]
[1]
|
||||
| Target | Bacterial Fatty acid synthetase I (Bact inhA) | INHA_MYCTU | [1] | ||
| Click to Show/Hide the Molecular Information and External Link(s) of This Drug | |||||
| Formula |
C6H7N3O
|
||||
| IsoSMILES |
C1=CN=CC=C1C(=O)NN
|
||||
| InChI |
1S/C6H7N3O/c7-9-6(10)5-1-3-8-4-2-5/h1-4H,7H2,(H,9,10)
|
||||
| InChIKey |
QRXWMOHMRWLFEY-UHFFFAOYSA-N
|
||||
| PubChem CID | |||||
| ChEBI ID | |||||
| TTD Drug ID | |||||
| VARIDT ID | |||||
| INTEDE ID | |||||
| DrugBank ID | |||||
Type(s) of Resistant Mechanism of This Drug
Drug Resistance Data Categorized by Their Corresponding Diseases
ICD-01: Infectious/parasitic diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Enoyl-[acyl-carrier-protein] reductase [NADH] (INHA) | [5] | |||
| Resistant Disease | Tuberculosis [ICD-11: 1B10.0] | |||
| Molecule Alteration | Mutation | . |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Mycobacterium tuberculosis H37Rv | 83332 | ||
| Mycobacterium tuberculosis isolates | 1773 | |||
| Experiment for Molecule Alteration |
qRT-PCR | |||
| Mechanism Description | Monoresistance to rifampicin and isoniazid was found in 11% (95% CI: 0.077-0.150; p, 0.087) and 8.5% (95% CI: 0.056-0.123; p, 0.692) of all the patients, respectively. Resistance to RIF and INH among newly diagnosed patients was 10.2% and 8.6%, while among previously treated patients, resistance to RIF and INH was 23.5% and 5.9% respectively. Furthermore, 4.9% of the samples from newly diagnosed with INH monoresistance, were found to have mutations in the InhA region while 8.6% had mutations in the katG region, a condition that can lead to phenotypic isoniazid drug resistance. | |||
| Key Molecule: Outer membrane protein A (OmpA) | [6] | |||
| Resistant Disease | Tuberculosis [ICD-11: 1B10.0] | |||
| Molecule Alteration | Expressiom | D96D |
||
| Experimental Note | Revealed Based on the Cell Line Data | |||
| In Vitro Model | Mycobacterium tuberculosis | 1773 | ||
| Experiment for Drug Resistance |
MIC assay | |||
| Mechanism Description | These results support the model that the roles of OmpA as a porin protein overexpressing in mycobacteria can increase the hydrophilic ability of the cell wall which can facilitate the streptomycin uptakes and increase the mycobacteria's sensitivity to aminoglycosides. | |||
|
|
||||
| Key Molecule: DNA-directed RNA polymerase subunit beta (RPOB) | [5] | |||
| Resistant Disease | Tuberculosis [ICD-11: 1B10.0] | |||
| Molecule Alteration | Mutation | . |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Mycobacterium tuberculosis H37Rv | 83332 | ||
| Mycobacterium tuberculosis isolates | 1773 | |||
| Experiment for Molecule Alteration |
qRT-PCR | |||
| Mechanism Description | Monoresistance to rifampicin and isoniazid was found in 11% (95% CI: 0.077-0.150; p, 0.087) and 8.5% (95% CI: 0.056-0.123; p, 0.692) of all the patients, respectively. Resistance to RIF and INH among newly diagnosed patients was 10.2% and 8.6%, while among previously treated patients, resistance to RIF and INH was 23.5% and 5.9% respectively. Furthermore, 4.9% of the samples from newly diagnosed with INH monoresistance, were found to have mutations in the InhA region while 8.6% had mutations in the katG region, a condition that can lead to phenotypic isoniazid drug resistance. | |||
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | |||||||||||||
|
|
|||||||||||||
| Key Molecule: Enoyl-[acyl-carrier-protein] reductase [NADH] (INHA) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S94A |
|||||||||||
| Wild Type Structure | Method: X-ray diffraction | Resolution: 1.40 Å | |||||||||||
| Mutant Type Structure | Method: X-ray diffraction | Resolution: 1.90 Å | |||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
-
M
T
T
G
G
L
L
L
L
D
D
G
G
K
K
R
R
10
|
I
I
L
L
V
V
S
S
G
G
I
I
I
I
T
T
D
D
S
S
20
|
S
S
I
I
A
A
F
F
H
H
I
I
A
A
R
R
V
V
A
A
30
|
Q
Q
E
E
Q
Q
G
G
A
A
Q
Q
L
L
V
V
L
L
T
T
40
|
G
G
F
F
D
D
R
R
L
L
R
R
L
L
I
I
Q
Q
R
R
50
|
I
I
T
T
D
D
R
R
L
L
P
P
A
A
K
K
A
A
P
P
60
|
L
L
L
L
E
E
L
L
D
D
V
V
Q
Q
N
N
E
E
E
E
70
|
H
H
L
L
A
A
S
S
L
L
A
A
G
G
R
R
V
V
T
T
80
|
E
E
A
A
I
I
G
G
A
A
G
G
N
N
K
K
L
L
D
D
90
|
G
G
V
V
V
V
H
H
S
A
I
I
G
G
F
F
M
M
P
P
100
|
Q
Q
T
T
G
G
M
M
G
G
I
I
N
N
P
P
F
F
F
F
110
|
D
D
A
A
P
P
Y
Y
A
A
D
D
V
V
S
S
K
K
G
G
120
|
I
I
H
H
I
I
S
S
A
A
Y
Y
S
S
Y
Y
A
A
S
S
130
|
M
M
A
A
K
K
A
A
L
L
L
L
P
P
I
I
M
M
N
N
140
|
P
P
G
G
G
G
S
S
I
I
V
V
G
G
M
M
D
D
F
F
150
|
D
D
P
P
S
S
R
R
A
A
M
M
P
P
A
A
Y
Y
N
N
160
|
W
W
M
M
T
T
V
V
A
A
K
K
S
S
A
A
L
L
E
E
170
|
S
S
V
V
N
N
R
R
F
F
V
V
A
A
R
R
E
E
A
A
180
|
G
G
K
K
Y
Y
G
G
V
V
R
R
S
S
N
N
L
L
V
V
190
|
A
A
A
A
G
G
P
P
I
I
R
R
T
T
L
L
A
A
M
M
200
|
S
S
A
A
I
I
V
V
G
G
G
G
A
A
L
L
G
G
E
E
210
|
E
E
A
A
G
G
A
A
Q
Q
I
I
Q
Q
L
L
L
L
E
E
220
|
E
E
G
G
W
W
D
D
Q
Q
R
R
A
A
P
P
I
I
G
G
230
|
W
W
N
N
M
M
K
K
D
D
A
A
T
T
P
P
V
V
A
A
240
|
K
K
T
T
V
V
C
C
A
A
L
L
L
L
S
S
D
D
W
W
250
|
L
L
P
P
A
A
T
T
T
T
G
G
D
D
I
I
I
I
Y
Y
260
|
A
A
D
D
G
G
G
G
A
A
H
H
T
T
Q
Q
L
L
L
L
|
|||||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Enoyl-[acyl-carrier-protein] reductase [NADH] (INHA) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.G141E |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Enoyl-[acyl-carrier-protein] reductase [NADH] (INHA) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.I194T |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
|
|
|||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S315T |
|||||||||||
| Wild Type Structure | Method: X-ray diffraction | Resolution: 2.00 Å | |||||||||||
| Mutant Type Structure | Method: X-ray diffraction | Resolution: 2.10 Å | |||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
M
M
P
P
E
E
Q
Q
H
H
P
P
P
P
I
I
T
T
10
|
E
E
T
T
T
T
T
T
G
G
A
A
A
A
S
S
N
N
G
G
20
|
C
C
P
P
V
V
V
V
G
G
H
H
M
M
K
K
Y
Y
P
P
30
|
V
V
E
E
G
G
G
G
G
G
N
N
Q
Q
D
D
W
W
W
W
40
|
P
P
N
N
R
R
L
L
N
N
L
L
K
K
V
V
L
L
H
H
50
|
Q
Q
N
N
P
P
A
A
V
V
A
A
D
D
P
P
M
M
G
G
60
|
A
A
A
A
F
F
D
D
Y
Y
A
A
A
A
E
E
V
V
A
A
70
|
T
T
I
I
D
D
V
V
D
D
A
A
L
L
T
T
R
R
D
D
80
|
I
I
E
E
E
E
V
V
M
M
T
T
T
T
S
S
Q
Q
P
P
90
|
W
W
W
W
P
P
A
A
D
D
Y
Y
G
G
H
H
Y
Y
G
G
100
|
P
P
L
L
F
F
I
I
R
R
M
M
A
A
W
W
H
H
A
A
110
|
A
A
G
G
T
T
Y
Y
R
R
I
I
H
H
D
D
G
G
R
R
120
|
G
G
G
G
A
A
G
G
G
G
G
G
M
M
Q
Q
R
R
F
F
130
|
A
A
P
P
L
L
N
N
S
S
W
W
P
P
D
D
N
N
A
A
140
|
S
S
L
L
D
D
K
K
A
A
R
R
R
R
L
L
L
L
W
W
150
|
P
P
V
V
K
K
K
K
K
K
Y
Y
G
G
K
K
K
K
L
L
160
|
S
S
W
W
A
A
D
D
L
L
I
I
V
V
F
F
A
A
G
G
170
|
N
N
C
C
A
A
L
L
E
E
S
S
M
M
G
G
F
F
K
K
180
|
T
T
F
F
G
G
F
F
G
G
F
F
G
G
R
R
V
V
D
D
190
|
Q
Q
W
W
E
E
P
P
D
D
E
E
V
V
Y
Y
W
W
G
G
200
|
K
K
E
E
A
A
T
T
W
W
L
L
G
G
D
D
E
E
R
R
210
|
Y
Y
S
S
G
G
K
K
R
R
D
D
L
L
E
E
N
N
P
P
220
|
L
L
A
A
A
A
V
V
Q
Q
M
M
G
G
L
L
I
I
Y
Y
230
|
V
V
N
N
P
P
E
E
G
G
P
P
N
N
G
G
N
N
P
P
240
|
D
D
P
P
M
M
A
A
A
A
A
A
V
V
D
D
I
I
R
R
250
|
E
E
T
T
F
F
R
R
R
R
M
M
A
A
M
M
N
N
D
D
260
|
V
V
E
E
T
T
A
A
A
A
L
L
I
I
V
V
G
G
G
G
270
|
H
H
T
T
F
F
G
G
K
K
T
T
H
H
G
G
A
A
G
G
280
|
P
P
A
A
D
D
L
L
V
V
G
G
P
P
E
E
P
P
E
E
290
|
A
A
A
A
P
P
L
L
E
E
Q
Q
M
M
G
G
L
L
G
G
300
|
W
W
K
K
S
S
S
S
Y
Y
G
G
T
T
G
G
T
T
G
G
310
|
K
K
D
D
A
A
I
I
T
T
S
T
G
G
I
I
E
E
V
V
320
|
V
V
W
W
T
T
N
N
T
T
P
P
T
T
K
K
W
W
D
D
330
|
N
N
S
S
F
F
L
L
E
E
I
I
L
L
Y
Y
G
G
Y
Y
340
|
E
E
W
W
E
E
L
L
T
T
K
K
S
S
P
P
A
A
G
G
350
|
A
A
W
W
Q
Q
Y
Y
T
T
A
A
K
K
D
D
G
G
A
A
360
|
G
G
A
A
G
G
T
T
I
I
P
P
D
D
P
P
F
F
G
G
370
|
G
G
P
P
G
G
R
R
S
S
P
P
T
T
M
M
L
L
A
A
380
|
T
T
D
D
L
L
S
S
L
L
R
R
V
V
D
D
P
P
I
I
390
|
Y
Y
E
E
R
R
I
I
T
T
R
R
R
R
W
W
L
L
E
E
400
|
H
H
P
P
E
E
E
E
L
L
A
A
D
D
E
E
F
F
A
A
410
|
K
K
A
A
W
W
Y
Y
K
K
L
L
I
I
H
H
R
R
D
D
420
|
M
M
G
G
P
P
V
V
A
A
R
R
Y
Y
L
L
G
G
P
P
430
|
L
L
V
V
P
P
K
K
Q
Q
T
T
L
L
L
L
W
W
Q
Q
440
|
D
D
P
P
V
V
P
P
A
A
V
V
S
S
H
H
D
D
L
L
450
|
V
V
G
G
E
E
A
A
E
E
I
I
A
A
S
S
L
L
K
K
460
|
S
S
Q
Q
I
I
R
R
A
A
S
S
G
G
L
L
T
T
V
V
470
|
S
S
Q
Q
L
L
V
V
S
S
T
T
A
A
W
W
A
A
A
A
480
|
A
A
S
S
S
S
F
F
R
R
G
G
S
S
D
D
K
K
R
R
490
|
G
G
G
G
A
A
N
N
G
G
G
G
R
R
I
I
R
R
L
L
500
|
Q
Q
P
P
Q
Q
V
V
G
G
W
W
E
E
V
V
N
N
D
D
510
|
P
P
D
D
G
G
D
D
L
L
R
R
K
K
V
V
I
I
R
R
520
|
T
T
L
L
E
E
E
E
I
I
Q
Q
E
E
S
S
F
F
N
N
530
|
S
S
A
A
A
A
P
P
G
G
N
N
I
I
K
K
V
V
S
S
540
|
F
F
A
A
D
D
L
L
V
V
V
V
L
L
G
G
G
G
C
C
550
|
A
A
A
A
I
I
E
E
K
K
A
A
A
A
K
K
A
A
A
A
560
|
G
G
H
H
N
N
I
I
T
T
V
V
P
P
F
F
T
T
P
P
570
|
G
G
R
R
T
T
D
D
A
A
S
S
Q
Q
E
E
Q
Q
T
T
580
|
D
D
V
V
E
E
S
S
F
F
A
A
V
V
L
L
E
E
P
P
590
|
K
K
A
A
D
D
G
G
F
F
R
R
N
N
Y
Y
L
L
G
G
600
|
K
K
G
G
N
N
P
P
L
L
P
P
A
A
E
E
Y
Y
M
M
610
|
L
L
L
L
D
D
K
K
A
A
N
N
L
L
L
L
T
T
L
L
620
|
S
S
A
A
P
P
E
E
M
M
T
T
V
V
L
L
V
V
G
G
630
|
G
G
L
L
R
R
V
V
L
L
G
G
A
A
N
N
Y
Y
K
K
640
|
R
R
L
L
P
P
L
L
G
G
V
V
F
F
T
T
E
E
A
A
650
|
S
S
E
E
S
S
L
L
T
T
N
N
D
D
F
F
F
F
V
V
660
|
N
N
L
L
L
L
D
D
M
M
G
G
I
I
T
T
W
W
E
E
670
|
P
P
S
S
P
P
A
A
D
D
D
D
G
G
T
T
Y
Y
Q
Q
680
|
G
G
K
K
D
D
G
G
S
S
G
G
K
K
V
V
K
K
W
W
690
|
T
T
G
G
S
S
R
R
V
V
D
D
L
L
V
V
F
F
G
G
700
|
S
S
N
N
S
S
E
E
L
L
R
R
A
A
L
L
V
V
E
E
710
|
V
V
Y
Y
G
G
A
A
D
D
D
D
A
A
Q
Q
P
P
K
K
720
|
F
F
V
V
Q
Q
D
D
F
F
V
V
A
A
A
A
W
W
D
D
730
|
K
K
V
V
M
M
N
N
L
L
D
D
R
R
F
F
D
D
V
V
740
|
R
R
|
|||||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S315T |
|||||||||||
| Wild Type Structure | Method: X-ray diffraction | Resolution: 2.00 Å | |||||||||||
| Mutant Type Structure | Method: X-ray diffraction | Resolution: 2.10 Å | |||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
M
M
P
P
E
E
Q
Q
H
H
P
P
P
P
I
I
T
T
10
|
E
E
T
T
T
T
T
T
G
G
A
A
A
A
S
S
N
N
G
G
20
|
C
C
P
P
V
V
V
V
G
G
H
H
M
M
K
K
Y
Y
P
P
30
|
V
V
E
E
G
G
G
G
G
G
N
N
Q
Q
D
D
W
W
W
W
40
|
P
P
N
N
R
R
L
L
N
N
L
L
K
K
V
V
L
L
H
H
50
|
Q
Q
N
N
P
P
A
A
V
V
A
A
D
D
P
P
M
M
G
G
60
|
A
A
A
A
F
F
D
D
Y
Y
A
A
A
A
E
E
V
V
A
A
70
|
T
T
I
I
D
D
V
V
D
D
A
A
L
L
T
T
R
R
D
D
80
|
I
I
E
E
E
E
V
V
M
M
T
T
T
T
S
S
Q
Q
P
P
90
|
W
W
W
W
P
P
A
A
D
D
Y
Y
G
G
H
H
Y
Y
G
G
100
|
P
P
L
L
F
F
I
I
R
R
M
M
A
A
W
W
H
H
A
A
110
|
A
A
G
G
T
T
Y
Y
R
R
I
I
H
H
D
D
G
G
R
R
120
|
G
G
G
G
A
A
G
G
G
G
G
G
M
M
Q
Q
R
R
F
F
130
|
A
A
P
P
L
L
N
N
S
S
W
W
P
P
D
D
N
N
A
A
140
|
S
S
L
L
D
D
K
K
A
A
R
R
R
R
L
L
L
L
W
W
150
|
P
P
V
V
K
K
K
K
K
K
Y
Y
G
G
K
K
K
K
L
L
160
|
S
S
W
W
A
A
D
D
L
L
I
I
V
V
F
F
A
A
G
G
170
|
N
N
C
C
A
A
L
L
E
E
S
S
M
M
G
G
F
F
K
K
180
|
T
T
F
F
G
G
F
F
G
G
F
F
G
G
R
R
V
V
D
D
190
|
Q
Q
W
W
E
E
P
P
D
D
E
E
V
V
Y
Y
W
W
G
G
200
|
K
K
E
E
A
A
T
T
W
W
L
L
G
G
D
D
E
E
R
R
210
|
Y
Y
S
S
G
G
K
K
R
R
D
D
L
L
E
E
N
N
P
P
220
|
L
L
A
A
A
A
V
V
Q
Q
M
M
G
G
L
L
I
I
Y
Y
230
|
V
V
N
N
P
P
E
E
G
G
P
P
N
N
G
G
N
N
P
P
240
|
D
D
P
P
M
M
A
A
A
A
A
A
V
V
D
D
I
I
R
R
250
|
E
E
T
T
F
F
R
R
R
R
M
M
A
A
M
M
N
N
D
D
260
|
V
V
E
E
T
T
A
A
A
A
L
L
I
I
V
V
G
G
G
G
270
|
H
H
T
T
F
F
G
G
K
K
T
T
H
H
G
G
A
A
G
G
280
|
P
P
A
A
D
D
L
L
V
V
G
G
P
P
E
E
P
P
E
E
290
|
A
A
A
A
P
P
L
L
E
E
Q
Q
M
M
G
G
L
L
G
G
300
|
W
W
K
K
S
S
S
S
Y
Y
G
G
T
T
G
G
T
T
G
G
310
|
K
K
D
D
A
A
I
I
T
T
S
T
G
G
I
I
E
E
V
V
320
|
V
V
W
W
T
T
N
N
T
T
P
P
T
T
K
K
W
W
D
D
330
|
N
N
S
S
F
F
L
L
E
E
I
I
L
L
Y
Y
G
G
Y
Y
340
|
E
E
W
W
E
E
L
L
T
T
K
K
S
S
P
P
A
A
G
G
350
|
A
A
W
W
Q
Q
Y
Y
T
T
A
A
K
K
D
D
G
G
A
A
360
|
G
G
A
A
G
G
T
T
I
I
P
P
D
D
P
P
F
F
G
G
370
|
G
G
P
P
G
G
R
R
S
S
P
P
T
T
M
M
L
L
A
A
380
|
T
T
D
D
L
L
S
S
L
L
R
R
V
V
D
D
P
P
I
I
390
|
Y
Y
E
E
R
R
I
I
T
T
R
R
R
R
W
W
L
L
E
E
400
|
H
H
P
P
E
E
E
E
L
L
A
A
D
D
E
E
F
F
A
A
410
|
K
K
A
A
W
W
Y
Y
K
K
L
L
I
I
H
H
R
R
D
D
420
|
M
M
G
G
P
P
V
V
A
A
R
R
Y
Y
L
L
G
G
P
P
430
|
L
L
V
V
P
P
K
K
Q
Q
T
T
L
L
L
L
W
W
Q
Q
440
|
D
D
P
P
V
V
P
P
A
A
V
V
S
S
H
H
D
D
L
L
450
|
V
V
G
G
E
E
A
A
E
E
I
I
A
A
S
S
L
L
K
K
460
|
S
S
Q
Q
I
I
R
R
A
A
S
S
G
G
L
L
T
T
V
V
470
|
S
S
Q
Q
L
L
V
V
S
S
T
T
A
A
W
W
A
A
A
A
480
|
A
A
S
S
S
S
F
F
R
R
G
G
S
S
D
D
K
K
R
R
490
|
G
G
G
G
A
A
N
N
G
G
G
G
R
R
I
I
R
R
L
L
500
|
Q
Q
P
P
Q
Q
V
V
G
G
W
W
E
E
V
V
N
N
D
D
510
|
P
P
D
D
G
G
D
D
L
L
R
R
K
K
V
V
I
I
R
R
520
|
T
T
L
L
E
E
E
E
I
I
Q
Q
E
E
S
S
F
F
N
N
530
|
S
S
A
A
A
A
P
P
G
G
N
N
I
I
K
K
V
V
S
S
540
|
F
F
A
A
D
D
L
L
V
V
V
V
L
L
G
G
G
G
C
C
550
|
A
A
A
A
I
I
E
E
K
K
A
A
A
A
K
K
A
A
A
A
560
|
G
G
H
H
N
N
I
I
T
T
V
V
P
P
F
F
T
T
P
P
570
|
G
G
R
R
T
T
D
D
A
A
S
S
Q
Q
E
E
Q
Q
T
T
580
|
D
D
V
V
E
E
S
S
F
F
A
A
V
V
L
L
E
E
P
P
590
|
K
K
A
A
D
D
G
G
F
F
R
R
N
N
Y
Y
L
L
G
G
600
|
K
K
G
G
N
N
P
P
L
L
P
P
A
A
E
E
Y
Y
M
M
610
|
L
L
L
L
D
D
K
K
A
A
N
N
L
L
L
L
T
T
L
L
620
|
S
S
A
A
P
P
E
E
M
M
T
T
V
V
L
L
V
V
G
G
630
|
G
G
L
L
R
R
V
V
L
L
G
G
A
A
N
N
Y
Y
K
K
640
|
R
R
L
L
P
P
L
L
G
G
V
V
F
F
T
T
E
E
A
A
650
|
S
S
E
E
S
S
L
L
T
T
N
N
D
D
F
F
F
F
V
V
660
|
N
N
L
L
L
L
D
D
M
M
G
G
I
I
T
T
W
W
E
E
670
|
P
P
S
S
P
P
A
A
D
D
D
D
G
G
T
T
Y
Y
Q
Q
680
|
G
G
K
K
D
D
G
G
S
S
G
G
K
K
V
V
K
K
W
W
690
|
T
T
G
G
S
S
R
R
V
V
D
D
L
L
V
V
F
F
G
G
700
|
S
S
N
N
S
S
E
E
L
L
R
R
A
A
L
L
V
V
E
E
710
|
V
V
Y
Y
G
G
A
A
D
D
D
D
A
A
Q
Q
P
P
K
K
720
|
F
F
V
V
Q
Q
D
D
F
F
V
V
A
A
A
A
W
W
D
D
730
|
K
K
V
V
M
M
N
N
L
L
D
D
R
R
F
F
D
D
V
V
740
|
R
R
|
|||||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S315T |
|||||||||||
| Wild Type Structure | Method: X-ray diffraction | Resolution: 2.00 Å | |||||||||||
| Mutant Type Structure | Method: X-ray diffraction | Resolution: 2.10 Å | |||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
M
M
P
P
E
E
Q
Q
H
H
P
P
P
P
I
I
T
T
10
|
E
E
T
T
T
T
T
T
G
G
A
A
A
A
S
S
N
N
G
G
20
|
C
C
P
P
V
V
V
V
G
G
H
H
M
M
K
K
Y
Y
P
P
30
|
V
V
E
E
G
G
G
G
G
G
N
N
Q
Q
D
D
W
W
W
W
40
|
P
P
N
N
R
R
L
L
N
N
L
L
K
K
V
V
L
L
H
H
50
|
Q
Q
N
N
P
P
A
A
V
V
A
A
D
D
P
P
M
M
G
G
60
|
A
A
A
A
F
F
D
D
Y
Y
A
A
A
A
E
E
V
V
A
A
70
|
T
T
I
I
D
D
V
V
D
D
A
A
L
L
T
T
R
R
D
D
80
|
I
I
E
E
E
E
V
V
M
M
T
T
T
T
S
S
Q
Q
P
P
90
|
W
W
W
W
P
P
A
A
D
D
Y
Y
G
G
H
H
Y
Y
G
G
100
|
P
P
L
L
F
F
I
I
R
R
M
M
A
A
W
W
H
H
A
A
110
|
A
A
G
G
T
T
Y
Y
R
R
I
I
H
H
D
D
G
G
R
R
120
|
G
G
G
G
A
A
G
G
G
G
G
G
M
M
Q
Q
R
R
F
F
130
|
A
A
P
P
L
L
N
N
S
S
W
W
P
P
D
D
N
N
A
A
140
|
S
S
L
L
D
D
K
K
A
A
R
R
R
R
L
L
L
L
W
W
150
|
P
P
V
V
K
K
K
K
K
K
Y
Y
G
G
K
K
K
K
L
L
160
|
S
S
W
W
A
A
D
D
L
L
I
I
V
V
F
F
A
A
G
G
170
|
N
N
C
C
A
A
L
L
E
E
S
S
M
M
G
G
F
F
K
K
180
|
T
T
F
F
G
G
F
F
G
G
F
F
G
G
R
R
V
V
D
D
190
|
Q
Q
W
W
E
E
P
P
D
D
E
E
V
V
Y
Y
W
W
G
G
200
|
K
K
E
E
A
A
T
T
W
W
L
L
G
G
D
D
E
E
R
R
210
|
Y
Y
S
S
G
G
K
K
R
R
D
D
L
L
E
E
N
N
P
P
220
|
L
L
A
A
A
A
V
V
Q
Q
M
M
G
G
L
L
I
I
Y
Y
230
|
V
V
N
N
P
P
E
E
G
G
P
P
N
N
G
G
N
N
P
P
240
|
D
D
P
P
M
M
A
A
A
A
A
A
V
V
D
D
I
I
R
R
250
|
E
E
T
T
F
F
R
R
R
R
M
M
A
A
M
M
N
N
D
D
260
|
V
V
E
E
T
T
A
A
A
A
L
L
I
I
V
V
G
G
G
G
270
|
H
H
T
T
F
F
G
G
K
K
T
T
H
H
G
G
A
A
G
G
280
|
P
P
A
A
D
D
L
L
V
V
G
G
P
P
E
E
P
P
E
E
290
|
A
A
A
A
P
P
L
L
E
E
Q
Q
M
M
G
G
L
L
G
G
300
|
W
W
K
K
S
S
S
S
Y
Y
G
G
T
T
G
G
T
T
G
G
310
|
K
K
D
D
A
A
I
I
T
T
S
T
G
G
I
I
E
E
V
V
320
|
V
V
W
W
T
T
N
N
T
T
P
P
T
T
K
K
W
W
D
D
330
|
N
N
S
S
F
F
L
L
E
E
I
I
L
L
Y
Y
G
G
Y
Y
340
|
E
E
W
W
E
E
L
L
T
T
K
K
S
S
P
P
A
A
G
G
350
|
A
A
W
W
Q
Q
Y
Y
T
T
A
A
K
K
D
D
G
G
A
A
360
|
G
G
A
A
G
G
T
T
I
I
P
P
D
D
P
P
F
F
G
G
370
|
G
G
P
P
G
G
R
R
S
S
P
P
T
T
M
M
L
L
A
A
380
|
T
T
D
D
L
L
S
S
L
L
R
R
V
V
D
D
P
P
I
I
390
|
Y
Y
E
E
R
R
I
I
T
T
R
R
R
R
W
W
L
L
E
E
400
|
H
H
P
P
E
E
E
E
L
L
A
A
D
D
E
E
F
F
A
A
410
|
K
K
A
A
W
W
Y
Y
K
K
L
L
I
I
H
H
R
R
D
D
420
|
M
M
G
G
P
P
V
V
A
A
R
R
Y
Y
L
L
G
G
P
P
430
|
L
L
V
V
P
P
K
K
Q
Q
T
T
L
L
L
L
W
W
Q
Q
440
|
D
D
P
P
V
V
P
P
A
A
V
V
S
S
H
H
D
D
L
L
450
|
V
V
G
G
E
E
A
A
E
E
I
I
A
A
S
S
L
L
K
K
460
|
S
S
Q
Q
I
I
R
R
A
A
S
S
G
G
L
L
T
T
V
V
470
|
S
S
Q
Q
L
L
V
V
S
S
T
T
A
A
W
W
A
A
A
A
480
|
A
A
S
S
S
S
F
F
R
R
G
G
S
S
D
D
K
K
R
R
490
|
G
G
G
G
A
A
N
N
G
G
G
G
R
R
I
I
R
R
L
L
500
|
Q
Q
P
P
Q
Q
V
V
G
G
W
W
E
E
V
V
N
N
D
D
510
|
P
P
D
D
G
G
D
D
L
L
R
R
K
K
V
V
I
I
R
R
520
|
T
T
L
L
E
E
E
E
I
I
Q
Q
E
E
S
S
F
F
N
N
530
|
S
S
A
A
A
A
P
P
G
G
N
N
I
I
K
K
V
V
S
S
540
|
F
F
A
A
D
D
L
L
V
V
V
V
L
L
G
G
G
G
C
C
550
|
A
A
A
A
I
I
E
E
K
K
A
A
A
A
K
K
A
A
A
A
560
|
G
G
H
H
N
N
I
I
T
T
V
V
P
P
F
F
T
T
P
P
570
|
G
G
R
R
T
T
D
D
A
A
S
S
Q
Q
E
E
Q
Q
T
T
580
|
D
D
V
V
E
E
S
S
F
F
A
A
V
V
L
L
E
E
P
P
590
|
K
K
A
A
D
D
G
G
F
F
R
R
N
N
Y
Y
L
L
G
G
600
|
K
K
G
G
N
N
P
P
L
L
P
P
A
A
E
E
Y
Y
M
M
610
|
L
L
L
L
D
D
K
K
A
A
N
N
L
L
L
L
T
T
L
L
620
|
S
S
A
A
P
P
E
E
M
M
T
T
V
V
L
L
V
V
G
G
630
|
G
G
L
L
R
R
V
V
L
L
G
G
A
A
N
N
Y
Y
K
K
640
|
R
R
L
L
P
P
L
L
G
G
V
V
F
F
T
T
E
E
A
A
650
|
S
S
E
E
S
S
L
L
T
T
N
N
D
D
F
F
F
F
V
V
660
|
N
N
L
L
L
L
D
D
M
M
G
G
I
I
T
T
W
W
E
E
670
|
P
P
S
S
P
P
A
A
D
D
D
D
G
G
T
T
Y
Y
Q
Q
680
|
G
G
K
K
D
D
G
G
S
S
G
G
K
K
V
V
K
K
W
W
690
|
T
T
G
G
S
S
R
R
V
V
D
D
L
L
V
V
F
F
G
G
700
|
S
S
N
N
S
S
E
E
L
L
R
R
A
A
L
L
V
V
E
E
710
|
V
V
Y
Y
G
G
A
A
D
D
D
D
A
A
Q
Q
P
P
K
K
720
|
F
F
V
V
Q
Q
D
D
F
F
V
V
A
A
A
A
W
W
D
D
730
|
K
K
V
V
M
M
N
N
L
L
D
D
R
R
F
F
D
D
V
V
740
|
R
R
|
|||||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S315T |
|||||||||||
| Wild Type Structure | Method: X-ray diffraction | Resolution: 2.00 Å | |||||||||||
| Mutant Type Structure | Method: X-ray diffraction | Resolution: 2.10 Å | |||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
M
M
P
P
E
E
Q
Q
H
H
P
P
P
P
I
I
T
T
10
|
E
E
T
T
T
T
T
T
G
G
A
A
A
A
S
S
N
N
G
G
20
|
C
C
P
P
V
V
V
V
G
G
H
H
M
M
K
K
Y
Y
P
P
30
|
V
V
E
E
G
G
G
G
G
G
N
N
Q
Q
D
D
W
W
W
W
40
|
P
P
N
N
R
R
L
L
N
N
L
L
K
K
V
V
L
L
H
H
50
|
Q
Q
N
N
P
P
A
A
V
V
A
A
D
D
P
P
M
M
G
G
60
|
A
A
A
A
F
F
D
D
Y
Y
A
A
A
A
E
E
V
V
A
A
70
|
T
T
I
I
D
D
V
V
D
D
A
A
L
L
T
T
R
R
D
D
80
|
I
I
E
E
E
E
V
V
M
M
T
T
T
T
S
S
Q
Q
P
P
90
|
W
W
W
W
P
P
A
A
D
D
Y
Y
G
G
H
H
Y
Y
G
G
100
|
P
P
L
L
F
F
I
I
R
R
M
M
A
A
W
W
H
H
A
A
110
|
A
A
G
G
T
T
Y
Y
R
R
I
I
H
H
D
D
G
G
R
R
120
|
G
G
G
G
A
A
G
G
G
G
G
G
M
M
Q
Q
R
R
F
F
130
|
A
A
P
P
L
L
N
N
S
S
W
W
P
P
D
D
N
N
A
A
140
|
S
S
L
L
D
D
K
K
A
A
R
R
R
R
L
L
L
L
W
W
150
|
P
P
V
V
K
K
K
K
K
K
Y
Y
G
G
K
K
K
K
L
L
160
|
S
S
W
W
A
A
D
D
L
L
I
I
V
V
F
F
A
A
G
G
170
|
N
N
C
C
A
A
L
L
E
E
S
S
M
M
G
G
F
F
K
K
180
|
T
T
F
F
G
G
F
F
G
G
F
F
G
G
R
R
V
V
D
D
190
|
Q
Q
W
W
E
E
P
P
D
D
E
E
V
V
Y
Y
W
W
G
G
200
|
K
K
E
E
A
A
T
T
W
W
L
L
G
G
D
D
E
E
R
R
210
|
Y
Y
S
S
G
G
K
K
R
R
D
D
L
L
E
E
N
N
P
P
220
|
L
L
A
A
A
A
V
V
Q
Q
M
M
G
G
L
L
I
I
Y
Y
230
|
V
V
N
N
P
P
E
E
G
G
P
P
N
N
G
G
N
N
P
P
240
|
D
D
P
P
M
M
A
A
A
A
A
A
V
V
D
D
I
I
R
R
250
|
E
E
T
T
F
F
R
R
R
R
M
M
A
A
M
M
N
N
D
D
260
|
V
V
E
E
T
T
A
A
A
A
L
L
I
I
V
V
G
G
G
G
270
|
H
H
T
T
F
F
G
G
K
K
T
T
H
H
G
G
A
A
G
G
280
|
P
P
A
A
D
D
L
L
V
V
G
G
P
P
E
E
P
P
E
E
290
|
A
A
A
A
P
P
L
L
E
E
Q
Q
M
M
G
G
L
L
G
G
300
|
W
W
K
K
S
S
S
S
Y
Y
G
G
T
T
G
G
T
T
G
G
310
|
K
K
D
D
A
A
I
I
T
T
S
T
G
G
I
I
E
E
V
V
320
|
V
V
W
W
T
T
N
N
T
T
P
P
T
T
K
K
W
W
D
D
330
|
N
N
S
S
F
F
L
L
E
E
I
I
L
L
Y
Y
G
G
Y
Y
340
|
E
E
W
W
E
E
L
L
T
T
K
K
S
S
P
P
A
A
G
G
350
|
A
A
W
W
Q
Q
Y
Y
T
T
A
A
K
K
D
D
G
G
A
A
360
|
G
G
A
A
G
G
T
T
I
I
P
P
D
D
P
P
F
F
G
G
370
|
G
G
P
P
G
G
R
R
S
S
P
P
T
T
M
M
L
L
A
A
380
|
T
T
D
D
L
L
S
S
L
L
R
R
V
V
D
D
P
P
I
I
390
|
Y
Y
E
E
R
R
I
I
T
T
R
R
R
R
W
W
L
L
E
E
400
|
H
H
P
P
E
E
E
E
L
L
A
A
D
D
E
E
F
F
A
A
410
|
K
K
A
A
W
W
Y
Y
K
K
L
L
I
I
H
H
R
R
D
D
420
|
M
M
G
G
P
P
V
V
A
A
R
R
Y
Y
L
L
G
G
P
P
430
|
L
L
V
V
P
P
K
K
Q
Q
T
T
L
L
L
L
W
W
Q
Q
440
|
D
D
P
P
V
V
P
P
A
A
V
V
S
S
H
H
D
D
L
L
450
|
V
V
G
G
E
E
A
A
E
E
I
I
A
A
S
S
L
L
K
K
460
|
S
S
Q
Q
I
I
R
R
A
A
S
S
G
G
L
L
T
T
V
V
470
|
S
S
Q
Q
L
L
V
V
S
S
T
T
A
A
W
W
A
A
A
A
480
|
A
A
S
S
S
S
F
F
R
R
G
G
S
S
D
D
K
K
R
R
490
|
G
G
G
G
A
A
N
N
G
G
G
G
R
R
I
I
R
R
L
L
500
|
Q
Q
P
P
Q
Q
V
V
G
G
W
W
E
E
V
V
N
N
D
D
510
|
P
P
D
D
G
G
D
D
L
L
R
R
K
K
V
V
I
I
R
R
520
|
T
T
L
L
E
E
E
E
I
I
Q
Q
E
E
S
S
F
F
N
N
530
|
S
S
A
A
A
A
P
P
G
G
N
N
I
I
K
K
V
V
S
S
540
|
F
F
A
A
D
D
L
L
V
V
V
V
L
L
G
G
G
G
C
C
550
|
A
A
A
A
I
I
E
E
K
K
A
A
A
A
K
K
A
A
A
A
560
|
G
G
H
H
N
N
I
I
T
T
V
V
P
P
F
F
T
T
P
P
570
|
G
G
R
R
T
T
D
D
A
A
S
S
Q
Q
E
E
Q
Q
T
T
580
|
D
D
V
V
E
E
S
S
F
F
A
A
V
V
L
L
E
E
P
P
590
|
K
K
A
A
D
D
G
G
F
F
R
R
N
N
Y
Y
L
L
G
G
600
|
K
K
G
G
N
N
P
P
L
L
P
P
A
A
E
E
Y
Y
M
M
610
|
L
L
L
L
D
D
K
K
A
A
N
N
L
L
L
L
T
T
L
L
620
|
S
S
A
A
P
P
E
E
M
M
T
T
V
V
L
L
V
V
G
G
630
|
G
G
L
L
R
R
V
V
L
L
G
G
A
A
N
N
Y
Y
K
K
640
|
R
R
L
L
P
P
L
L
G
G
V
V
F
F
T
T
E
E
A
A
650
|
S
S
E
E
S
S
L
L
T
T
N
N
D
D
F
F
F
F
V
V
660
|
N
N
L
L
L
L
D
D
M
M
G
G
I
I
T
T
W
W
E
E
670
|
P
P
S
S
P
P
A
A
D
D
D
D
G
G
T
T
Y
Y
Q
Q
680
|
G
G
K
K
D
D
G
G
S
S
G
G
K
K
V
V
K
K
W
W
690
|
T
T
G
G
S
S
R
R
V
V
D
D
L
L
V
V
F
F
G
G
700
|
S
S
N
N
S
S
E
E
L
L
R
R
A
A
L
L
V
V
E
E
710
|
V
V
Y
Y
G
G
A
A
D
D
D
D
A
A
Q
Q
P
P
K
K
720
|
F
F
V
V
Q
Q
D
D
F
F
V
V
A
A
A
A
W
W
D
D
730
|
K
K
V
V
M
M
N
N
L
L
D
D
R
R
F
F
D
D
V
V
740
|
R
R
|
|||||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S315T |
|||||||||||
| Wild Type Structure | Method: X-ray diffraction | Resolution: 2.00 Å | |||||||||||
| Mutant Type Structure | Method: X-ray diffraction | Resolution: 2.10 Å | |||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
M
M
P
P
E
E
Q
Q
H
H
P
P
P
P
I
I
T
T
10
|
E
E
T
T
T
T
T
T
G
G
A
A
A
A
S
S
N
N
G
G
20
|
C
C
P
P
V
V
V
V
G
G
H
H
M
M
K
K
Y
Y
P
P
30
|
V
V
E
E
G
G
G
G
G
G
N
N
Q
Q
D
D
W
W
W
W
40
|
P
P
N
N
R
R
L
L
N
N
L
L
K
K
V
V
L
L
H
H
50
|
Q
Q
N
N
P
P
A
A
V
V
A
A
D
D
P
P
M
M
G
G
60
|
A
A
A
A
F
F
D
D
Y
Y
A
A
A
A
E
E
V
V
A
A
70
|
T
T
I
I
D
D
V
V
D
D
A
A
L
L
T
T
R
R
D
D
80
|
I
I
E
E
E
E
V
V
M
M
T
T
T
T
S
S
Q
Q
P
P
90
|
W
W
W
W
P
P
A
A
D
D
Y
Y
G
G
H
H
Y
Y
G
G
100
|
P
P
L
L
F
F
I
I
R
R
M
M
A
A
W
W
H
H
A
A
110
|
A
A
G
G
T
T
Y
Y
R
R
I
I
H
H
D
D
G
G
R
R
120
|
G
G
G
G
A
A
G
G
G
G
G
G
M
M
Q
Q
R
R
F
F
130
|
A
A
P
P
L
L
N
N
S
S
W
W
P
P
D
D
N
N
A
A
140
|
S
S
L
L
D
D
K
K
A
A
R
R
R
R
L
L
L
L
W
W
150
|
P
P
V
V
K
K
K
K
K
K
Y
Y
G
G
K
K
K
K
L
L
160
|
S
S
W
W
A
A
D
D
L
L
I
I
V
V
F
F
A
A
G
G
170
|
N
N
C
C
A
A
L
L
E
E
S
S
M
M
G
G
F
F
K
K
180
|
T
T
F
F
G
G
F
F
G
G
F
F
G
G
R
R
V
V
D
D
190
|
Q
Q
W
W
E
E
P
P
D
D
E
E
V
V
Y
Y
W
W
G
G
200
|
K
K
E
E
A
A
T
T
W
W
L
L
G
G
D
D
E
E
R
R
210
|
Y
Y
S
S
G
G
K
K
R
R
D
D
L
L
E
E
N
N
P
P
220
|
L
L
A
A
A
A
V
V
Q
Q
M
M
G
G
L
L
I
I
Y
Y
230
|
V
V
N
N
P
P
E
E
G
G
P
P
N
N
G
G
N
N
P
P
240
|
D
D
P
P
M
M
A
A
A
A
A
A
V
V
D
D
I
I
R
R
250
|
E
E
T
T
F
F
R
R
R
R
M
M
A
A
M
M
N
N
D
D
260
|
V
V
E
E
T
T
A
A
A
A
L
L
I
I
V
V
G
G
G
G
270
|
H
H
T
T
F
F
G
G
K
K
T
T
H
H
G
G
A
A
G
G
280
|
P
P
A
A
D
D
L
L
V
V
G
G
P
P
E
E
P
P
E
E
290
|
A
A
A
A
P
P
L
L
E
E
Q
Q
M
M
G
G
L
L
G
G
300
|
W
W
K
K
S
S
S
S
Y
Y
G
G
T
T
G
G
T
T
G
G
310
|
K
K
D
D
A
A
I
I
T
T
S
T
G
G
I
I
E
E
V
V
320
|
V
V
W
W
T
T
N
N
T
T
P
P
T
T
K
K
W
W
D
D
330
|
N
N
S
S
F
F
L
L
E
E
I
I
L
L
Y
Y
G
G
Y
Y
340
|
E
E
W
W
E
E
L
L
T
T
K
K
S
S
P
P
A
A
G
G
350
|
A
A
W
W
Q
Q
Y
Y
T
T
A
A
K
K
D
D
G
G
A
A
360
|
G
G
A
A
G
G
T
T
I
I
P
P
D
D
P
P
F
F
G
G
370
|
G
G
P
P
G
G
R
R
S
S
P
P
T
T
M
M
L
L
A
A
380
|
T
T
D
D
L
L
S
S
L
L
R
R
V
V
D
D
P
P
I
I
390
|
Y
Y
E
E
R
R
I
I
T
T
R
R
R
R
W
W
L
L
E
E
400
|
H
H
P
P
E
E
E
E
L
L
A
A
D
D
E
E
F
F
A
A
410
|
K
K
A
A
W
W
Y
Y
K
K
L
L
I
I
H
H
R
R
D
D
420
|
M
M
G
G
P
P
V
V
A
A
R
R
Y
Y
L
L
G
G
P
P
430
|
L
L
V
V
P
P
K
K
Q
Q
T
T
L
L
L
L
W
W
Q
Q
440
|
D
D
P
P
V
V
P
P
A
A
V
V
S
S
H
H
D
D
L
L
450
|
V
V
G
G
E
E
A
A
E
E
I
I
A
A
S
S
L
L
K
K
460
|
S
S
Q
Q
I
I
R
R
A
A
S
S
G
G
L
L
T
T
V
V
470
|
S
S
Q
Q
L
L
V
V
S
S
T
T
A
A
W
W
A
A
A
A
480
|
A
A
S
S
S
S
F
F
R
R
G
G
S
S
D
D
K
K
R
R
490
|
G
G
G
G
A
A
N
N
G
G
G
G
R
R
I
I
R
R
L
L
500
|
Q
Q
P
P
Q
Q
V
V
G
G
W
W
E
E
V
V
N
N
D
D
510
|
P
P
D
D
G
G
D
D
L
L
R
R
K
K
V
V
I
I
R
R
520
|
T
T
L
L
E
E
E
E
I
I
Q
Q
E
E
S
S
F
F
N
N
530
|
S
S
A
A
A
A
P
P
G
G
N
N
I
I
K
K
V
V
S
S
540
|
F
F
A
A
D
D
L
L
V
V
V
V
L
L
G
G
G
G
C
C
550
|
A
A
A
A
I
I
E
E
K
K
A
A
A
A
K
K
A
A
A
A
560
|
G
G
H
H
N
N
I
I
T
T
V
V
P
P
F
F
T
T
P
P
570
|
G
G
R
R
T
T
D
D
A
A
S
S
Q
Q
E
E
Q
Q
T
T
580
|
D
D
V
V
E
E
S
S
F
F
A
A
V
V
L
L
E
E
P
P
590
|
K
K
A
A
D
D
G
G
F
F
R
R
N
N
Y
Y
L
L
G
G
600
|
K
K
G
G
N
N
P
P
L
L
P
P
A
A
E
E
Y
Y
M
M
610
|
L
L
L
L
D
D
K
K
A
A
N
N
L
L
L
L
T
T
L
L
620
|
S
S
A
A
P
P
E
E
M
M
T
T
V
V
L
L
V
V
G
G
630
|
G
G
L
L
R
R
V
V
L
L
G
G
A
A
N
N
Y
Y
K
K
640
|
R
R
L
L
P
P
L
L
G
G
V
V
F
F
T
T
E
E
A
A
650
|
S
S
E
E
S
S
L
L
T
T
N
N
D
D
F
F
F
F
V
V
660
|
N
N
L
L
L
L
D
D
M
M
G
G
I
I
T
T
W
W
E
E
670
|
P
P
S
S
P
P
A
A
D
D
D
D
G
G
T
T
Y
Y
Q
Q
680
|
G
G
K
K
D
D
G
G
S
S
G
G
K
K
V
V
K
K
W
W
690
|
T
T
G
G
S
S
R
R
V
V
D
D
L
L
V
V
F
F
G
G
700
|
S
S
N
N
S
S
E
E
L
L
R
R
A
A
L
L
V
V
E
E
710
|
V
V
Y
Y
G
G
A
A
D
D
D
D
A
A
Q
Q
P
P
K
K
720
|
F
F
V
V
Q
Q
D
D
F
F
V
V
A
A
A
A
W
W
D
D
730
|
K
K
V
V
M
M
N
N
L
L
D
D
R
R
F
F
D
D
V
V
740
|
R
R
|
|||||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S315T |
|||||||||||
| Wild Type Structure | Method: X-ray diffraction | Resolution: 2.00 Å | |||||||||||
| Mutant Type Structure | Method: X-ray diffraction | Resolution: 2.10 Å | |||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
M
M
P
P
E
E
Q
Q
H
H
P
P
P
P
I
I
T
T
10
|
E
E
T
T
T
T
T
T
G
G
A
A
A
A
S
S
N
N
G
G
20
|
C
C
P
P
V
V
V
V
G
G
H
H
M
M
K
K
Y
Y
P
P
30
|
V
V
E
E
G
G
G
G
G
G
N
N
Q
Q
D
D
W
W
W
W
40
|
P
P
N
N
R
R
L
L
N
N
L
L
K
K
V
V
L
L
H
H
50
|
Q
Q
N
N
P
P
A
A
V
V
A
A
D
D
P
P
M
M
G
G
60
|
A
A
A
A
F
F
D
D
Y
Y
A
A
A
A
E
E
V
V
A
A
70
|
T
T
I
I
D
D
V
V
D
D
A
A
L
L
T
T
R
R
D
D
80
|
I
I
E
E
E
E
V
V
M
M
T
T
T
T
S
S
Q
Q
P
P
90
|
W
W
W
W
P
P
A
A
D
D
Y
Y
G
G
H
H
Y
Y
G
G
100
|
P
P
L
L
F
F
I
I
R
R
M
M
A
A
W
W
H
H
A
A
110
|
A
A
G
G
T
T
Y
Y
R
R
I
I
H
H
D
D
G
G
R
R
120
|
G
G
G
G
A
A
G
G
G
G
G
G
M
M
Q
Q
R
R
F
F
130
|
A
A
P
P
L
L
N
N
S
S
W
W
P
P
D
D
N
N
A
A
140
|
S
S
L
L
D
D
K
K
A
A
R
R
R
R
L
L
L
L
W
W
150
|
P
P
V
V
K
K
K
K
K
K
Y
Y
G
G
K
K
K
K
L
L
160
|
S
S
W
W
A
A
D
D
L
L
I
I
V
V
F
F
A
A
G
G
170
|
N
N
C
C
A
A
L
L
E
E
S
S
M
M
G
G
F
F
K
K
180
|
T
T
F
F
G
G
F
F
G
G
F
F
G
G
R
R
V
V
D
D
190
|
Q
Q
W
W
E
E
P
P
D
D
E
E
V
V
Y
Y
W
W
G
G
200
|
K
K
E
E
A
A
T
T
W
W
L
L
G
G
D
D
E
E
R
R
210
|
Y
Y
S
S
G
G
K
K
R
R
D
D
L
L
E
E
N
N
P
P
220
|
L
L
A
A
A
A
V
V
Q
Q
M
M
G
G
L
L
I
I
Y
Y
230
|
V
V
N
N
P
P
E
E
G
G
P
P
N
N
G
G
N
N
P
P
240
|
D
D
P
P
M
M
A
A
A
A
A
A
V
V
D
D
I
I
R
R
250
|
E
E
T
T
F
F
R
R
R
R
M
M
A
A
M
M
N
N
D
D
260
|
V
V
E
E
T
T
A
A
A
A
L
L
I
I
V
V
G
G
G
G
270
|
H
H
T
T
F
F
G
G
K
K
T
T
H
H
G
G
A
A
G
G
280
|
P
P
A
A
D
D
L
L
V
V
G
G
P
P
E
E
P
P
E
E
290
|
A
A
A
A
P
P
L
L
E
E
Q
Q
M
M
G
G
L
L
G
G
300
|
W
W
K
K
S
S
S
S
Y
Y
G
G
T
T
G
G
T
T
G
G
310
|
K
K
D
D
A
A
I
I
T
T
S
T
G
G
I
I
E
E
V
V
320
|
V
V
W
W
T
T
N
N
T
T
P
P
T
T
K
K
W
W
D
D
330
|
N
N
S
S
F
F
L
L
E
E
I
I
L
L
Y
Y
G
G
Y
Y
340
|
E
E
W
W
E
E
L
L
T
T
K
K
S
S
P
P
A
A
G
G
350
|
A
A
W
W
Q
Q
Y
Y
T
T
A
A
K
K
D
D
G
G
A
A
360
|
G
G
A
A
G
G
T
T
I
I
P
P
D
D
P
P
F
F
G
G
370
|
G
G
P
P
G
G
R
R
S
S
P
P
T
T
M
M
L
L
A
A
380
|
T
T
D
D
L
L
S
S
L
L
R
R
V
V
D
D
P
P
I
I
390
|
Y
Y
E
E
R
R
I
I
T
T
R
R
R
R
W
W
L
L
E
E
400
|
H
H
P
P
E
E
E
E
L
L
A
A
D
D
E
E
F
F
A
A
410
|
K
K
A
A
W
W
Y
Y
K
K
L
L
I
I
H
H
R
R
D
D
420
|
M
M
G
G
P
P
V
V
A
A
R
R
Y
Y
L
L
G
G
P
P
430
|
L
L
V
V
P
P
K
K
Q
Q
T
T
L
L
L
L
W
W
Q
Q
440
|
D
D
P
P
V
V
P
P
A
A
V
V
S
S
H
H
D
D
L
L
450
|
V
V
G
G
E
E
A
A
E
E
I
I
A
A
S
S
L
L
K
K
460
|
S
S
Q
Q
I
I
R
R
A
A
S
S
G
G
L
L
T
T
V
V
470
|
S
S
Q
Q
L
L
V
V
S
S
T
T
A
A
W
W
A
A
A
A
480
|
A
A
S
S
S
S
F
F
R
R
G
G
S
S
D
D
K
K
R
R
490
|
G
G
G
G
A
A
N
N
G
G
G
G
R
R
I
I
R
R
L
L
500
|
Q
Q
P
P
Q
Q
V
V
G
G
W
W
E
E
V
V
N
N
D
D
510
|
P
P
D
D
G
G
D
D
L
L
R
R
K
K
V
V
I
I
R
R
520
|
T
T
L
L
E
E
E
E
I
I
Q
Q
E
E
S
S
F
F
N
N
530
|
S
S
A
A
A
A
P
P
G
G
N
N
I
I
K
K
V
V
S
S
540
|
F
F
A
A
D
D
L
L
V
V
V
V
L
L
G
G
G
G
C
C
550
|
A
A
A
A
I
I
E
E
K
K
A
A
A
A
K
K
A
A
A
A
560
|
G
G
H
H
N
N
I
I
T
T
V
V
P
P
F
F
T
T
P
P
570
|
G
G
R
R
T
T
D
D
A
A
S
S
Q
Q
E
E
Q
Q
T
T
580
|
D
D
V
V
E
E
S
S
F
F
A
A
V
V
L
L
E
E
P
P
590
|
K
K
A
A
D
D
G
G
F
F
R
R
N
N
Y
Y
L
L
G
G
600
|
K
K
G
G
N
N
P
P
L
L
P
P
A
A
E
E
Y
Y
M
M
610
|
L
L
L
L
D
D
K
K
A
A
N
N
L
L
L
L
T
T
L
L
620
|
S
S
A
A
P
P
E
E
M
M
T
T
V
V
L
L
V
V
G
G
630
|
G
G
L
L
R
R
V
V
L
L
G
G
A
A
N
N
Y
Y
K
K
640
|
R
R
L
L
P
P
L
L
G
G
V
V
F
F
T
T
E
E
A
A
650
|
S
S
E
E
S
S
L
L
T
T
N
N
D
D
F
F
F
F
V
V
660
|
N
N
L
L
L
L
D
D
M
M
G
G
I
I
T
T
W
W
E
E
670
|
P
P
S
S
P
P
A
A
D
D
D
D
G
G
T
T
Y
Y
Q
Q
680
|
G
G
K
K
D
D
G
G
S
S
G
G
K
K
V
V
K
K
W
W
690
|
T
T
G
G
S
S
R
R
V
V
D
D
L
L
V
V
F
F
G
G
700
|
S
S
N
N
S
S
E
E
L
L
R
R
A
A
L
L
V
V
E
E
710
|
V
V
Y
Y
G
G
A
A
D
D
D
D
A
A
Q
Q
P
P
K
K
720
|
F
F
V
V
Q
Q
D
D
F
F
V
V
A
A
A
A
W
W
D
D
730
|
K
K
V
V
M
M
N
N
L
L
D
D
R
R
F
F
D
D
V
V
740
|
R
R
|
|||||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S315T |
|||||||||||
| Wild Type Structure | Method: X-ray diffraction | Resolution: 2.00 Å | |||||||||||
| Mutant Type Structure | Method: X-ray diffraction | Resolution: 2.10 Å | |||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
M
M
P
P
E
E
Q
Q
H
H
P
P
P
P
I
I
T
T
10
|
E
E
T
T
T
T
T
T
G
G
A
A
A
A
S
S
N
N
G
G
20
|
C
C
P
P
V
V
V
V
G
G
H
H
M
M
K
K
Y
Y
P
P
30
|
V
V
E
E
G
G
G
G
G
G
N
N
Q
Q
D
D
W
W
W
W
40
|
P
P
N
N
R
R
L
L
N
N
L
L
K
K
V
V
L
L
H
H
50
|
Q
Q
N
N
P
P
A
A
V
V
A
A
D
D
P
P
M
M
G
G
60
|
A
A
A
A
F
F
D
D
Y
Y
A
A
A
A
E
E
V
V
A
A
70
|
T
T
I
I
D
D
V
V
D
D
A
A
L
L
T
T
R
R
D
D
80
|
I
I
E
E
E
E
V
V
M
M
T
T
T
T
S
S
Q
Q
P
P
90
|
W
W
W
W
P
P
A
A
D
D
Y
Y
G
G
H
H
Y
Y
G
G
100
|
P
P
L
L
F
F
I
I
R
R
M
M
A
A
W
W
H
H
A
A
110
|
A
A
G
G
T
T
Y
Y
R
R
I
I
H
H
D
D
G
G
R
R
120
|
G
G
G
G
A
A
G
G
G
G
G
G
M
M
Q
Q
R
R
F
F
130
|
A
A
P
P
L
L
N
N
S
S
W
W
P
P
D
D
N
N
A
A
140
|
S
S
L
L
D
D
K
K
A
A
R
R
R
R
L
L
L
L
W
W
150
|
P
P
V
V
K
K
K
K
K
K
Y
Y
G
G
K
K
K
K
L
L
160
|
S
S
W
W
A
A
D
D
L
L
I
I
V
V
F
F
A
A
G
G
170
|
N
N
C
C
A
A
L
L
E
E
S
S
M
M
G
G
F
F
K
K
180
|
T
T
F
F
G
G
F
F
G
G
F
F
G
G
R
R
V
V
D
D
190
|
Q
Q
W
W
E
E
P
P
D
D
E
E
V
V
Y
Y
W
W
G
G
200
|
K
K
E
E
A
A
T
T
W
W
L
L
G
G
D
D
E
E
R
R
210
|
Y
Y
S
S
G
G
K
K
R
R
D
D
L
L
E
E
N
N
P
P
220
|
L
L
A
A
A
A
V
V
Q
Q
M
M
G
G
L
L
I
I
Y
Y
230
|
V
V
N
N
P
P
E
E
G
G
P
P
N
N
G
G
N
N
P
P
240
|
D
D
P
P
M
M
A
A
A
A
A
A
V
V
D
D
I
I
R
R
250
|
E
E
T
T
F
F
R
R
R
R
M
M
A
A
M
M
N
N
D
D
260
|
V
V
E
E
T
T
A
A
A
A
L
L
I
I
V
V
G
G
G
G
270
|
H
H
T
T
F
F
G
G
K
K
T
T
H
H
G
G
A
A
G
G
280
|
P
P
A
A
D
D
L
L
V
V
G
G
P
P
E
E
P
P
E
E
290
|
A
A
A
A
P
P
L
L
E
E
Q
Q
M
M
G
G
L
L
G
G
300
|
W
W
K
K
S
S
S
S
Y
Y
G
G
T
T
G
G
T
T
G
G
310
|
K
K
D
D
A
A
I
I
T
T
S
T
G
G
I
I
E
E
V
V
320
|
V
V
W
W
T
T
N
N
T
T
P
P
T
T
K
K
W
W
D
D
330
|
N
N
S
S
F
F
L
L
E
E
I
I
L
L
Y
Y
G
G
Y
Y
340
|
E
E
W
W
E
E
L
L
T
T
K
K
S
S
P
P
A
A
G
G
350
|
A
A
W
W
Q
Q
Y
Y
T
T
A
A
K
K
D
D
G
G
A
A
360
|
G
G
A
A
G
G
T
T
I
I
P
P
D
D
P
P
F
F
G
G
370
|
G
G
P
P
G
G
R
R
S
S
P
P
T
T
M
M
L
L
A
A
380
|
T
T
D
D
L
L
S
S
L
L
R
R
V
V
D
D
P
P
I
I
390
|
Y
Y
E
E
R
R
I
I
T
T
R
R
R
R
W
W
L
L
E
E
400
|
H
H
P
P
E
E
E
E
L
L
A
A
D
D
E
E
F
F
A
A
410
|
K
K
A
A
W
W
Y
Y
K
K
L
L
I
I
H
H
R
R
D
D
420
|
M
M
G
G
P
P
V
V
A
A
R
R
Y
Y
L
L
G
G
P
P
430
|
L
L
V
V
P
P
K
K
Q
Q
T
T
L
L
L
L
W
W
Q
Q
440
|
D
D
P
P
V
V
P
P
A
A
V
V
S
S
H
H
D
D
L
L
450
|
V
V
G
G
E
E
A
A
E
E
I
I
A
A
S
S
L
L
K
K
460
|
S
S
Q
Q
I
I
R
R
A
A
S
S
G
G
L
L
T
T
V
V
470
|
S
S
Q
Q
L
L
V
V
S
S
T
T
A
A
W
W
A
A
A
A
480
|
A
A
S
S
S
S
F
F
R
R
G
G
S
S
D
D
K
K
R
R
490
|
G
G
G
G
A
A
N
N
G
G
G
G
R
R
I
I
R
R
L
L
500
|
Q
Q
P
P
Q
Q
V
V
G
G
W
W
E
E
V
V
N
N
D
D
510
|
P
P
D
D
G
G
D
D
L
L
R
R
K
K
V
V
I
I
R
R
520
|
T
T
L
L
E
E
E
E
I
I
Q
Q
E
E
S
S
F
F
N
N
530
|
S
S
A
A
A
A
P
P
G
G
N
N
I
I
K
K
V
V
S
S
540
|
F
F
A
A
D
D
L
L
V
V
V
V
L
L
G
G
G
G
C
C
550
|
A
A
A
A
I
I
E
E
K
K
A
A
A
A
K
K
A
A
A
A
560
|
G
G
H
H
N
N
I
I
T
T
V
V
P
P
F
F
T
T
P
P
570
|
G
G
R
R
T
T
D
D
A
A
S
S
Q
Q
E
E
Q
Q
T
T
580
|
D
D
V
V
E
E
S
S
F
F
A
A
V
V
L
L
E
E
P
P
590
|
K
K
A
A
D
D
G
G
F
F
R
R
N
N
Y
Y
L
L
G
G
600
|
K
K
G
G
N
N
P
P
L
L
P
P
A
A
E
E
Y
Y
M
M
610
|
L
L
L
L
D
D
K
K
A
A
N
N
L
L
L
L
T
T
L
L
620
|
S
S
A
A
P
P
E
E
M
M
T
T
V
V
L
L
V
V
G
G
630
|
G
G
L
L
R
R
V
V
L
L
G
G
A
A
N
N
Y
Y
K
K
640
|
R
R
L
L
P
P
L
L
G
G
V
V
F
F
T
T
E
E
A
A
650
|
S
S
E
E
S
S
L
L
T
T
N
N
D
D
F
F
F
F
V
V
660
|
N
N
L
L
L
L
D
D
M
M
G
G
I
I
T
T
W
W
E
E
670
|
P
P
S
S
P
P
A
A
D
D
D
D
G
G
T
T
Y
Y
Q
Q
680
|
G
G
K
K
D
D
G
G
S
S
G
G
K
K
V
V
K
K
W
W
690
|
T
T
G
G
S
S
R
R
V
V
D
D
L
L
V
V
F
F
G
G
700
|
S
S
N
N
S
S
E
E
L
L
R
R
A
A
L
L
V
V
E
E
710
|
V
V
Y
Y
G
G
A
A
D
D
D
D
A
A
Q
Q
P
P
K
K
720
|
F
F
V
V
Q
Q
D
D
F
F
V
V
A
A
A
A
W
W
D
D
730
|
K
K
V
V
M
M
N
N
L
L
D
D
R
R
F
F
D
D
V
V
740
|
R
R
|
|||||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S315N |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.A312P |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.A264V |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.N660D |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.L147P |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.C20R |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.T308P |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.T275A |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.D142G |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S211G |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.M126I |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.W91R |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S315G |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.G490S |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.V581G |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.A110V |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.G466R |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.G279V |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.L436P |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.N508D |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.P92S |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.G125S |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.Q127P |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.V431A |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.G490S |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.Q461P |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.E607A |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.H417Q |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.G111S |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.G33V |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Missense mutation | p.W191R |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
|
|
|||||||||||||
| Key Molecule: P-type ATPase zinc transporter Rv3270 | [3] | ||||||||||||
| Resistant Disease | Bone infection [ICD-11: 1B2Z.9] | ||||||||||||
| Molecule Alteration | Expression | Up-regulation |
|||||||||||
| Experimental Note | Revealed Based on the Cell Line Data | ||||||||||||
| In Vitro Model | E. coli XL1-Blue | 562 | |||||||||||
| E. coli CS109 | 562 | ||||||||||||
| M. smegmatis MC2 161 | 1772 | ||||||||||||
| Experiment for Molecule Alteration |
Gene expression analysis | ||||||||||||
| Experiment for Drug Resistance |
Antimicrobial susceptibility assay; Intracellular drug accumulation activity assay | ||||||||||||
| Mechanism Description | Metal homeostasis is maintained by the uptake, storage and efflux of metal ions that are necessary for the survival of the bacterium. Homeostasis is mostly regulated by a group of transporters categorized as ABC transporters and P-type ATPases. On the other hand, efflux pumps often play a role in drug-metal cross-resistance. Here, with the help of antibiotic sensitivity, antibiotic/dye accumulation and semi-quantitative biofilm formation assessments we report the ability of Rv3270, a P-type ATPase known for its role in combating Mn2+ and Zn2+ metal ion toxicity in Mycobacterium tuberculosis, in influencing the extrusion of multiple structurally unrelated drugs and enhancing the biofilm formation of Escherichia coli and Mycobacterium smegmatis. Overexpression of Rv3270 increased the tolerance of host cells to norfloxacin, ofloxacin, sparfloxacin, ampicillin, oxacillin, amikacin and isoniazid. A significantly lower accumulation of norfloxacin, ethidium bromide, bocillin FL and levofloxacin in cells harbouring Rv3270 as compared to host cells indicated its role in enhancing efflux activity. Although over-expression of Rv3270 did not alter the susceptibility levels of levofloxacin, rifampicin and apramycin, the presence of a sub-inhibitory concentration of Zn2+ resulted in low-level tolerance towards these drugs. Of note, the expression of Rv3270 enhanced the biofilm-forming ability of the host cells strengthening its role in antimicrobial resistance. Therefore, the study indicated that the over-expression of Rv3270 enhances the drug efflux activity of the micro-organism where zinc might facilitate drug-metal cross-resistance for some antibiotics. | ||||||||||||
|
|
|||||||||||||
| Key Molecule: NAD-dependent protein deacylase Sir2 (SIR2) | [1] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Expression | Up-regulation |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Escherichia coli | 668369 | |||||||||||
| Mycobacterium smegmatis mc2155 | 246196 | ||||||||||||
| Experiment for Molecule Alteration |
Quantitative Real-Time PCR | ||||||||||||
| Experiment for Drug Resistance |
Colony forming units determination assay | ||||||||||||
| Mechanism Description | MSMEG_5175 regulates diverse cellular processes resulting in an increase in INH resistance in mycobacteria. Overexpression of MSMEG_5175 results in up-regulation of 34 proteins and down-regulation of 72 proteins, which involve in diverse cellular processes including metabolic activation, transcription and translation, antioxidant, and DNA repair. Down-regulation of catalase peroxidase (katG) expression in both mRNA and protein levels were observed in mc(2)155-MS5175 strain, suggesting that a decrease in cellular NAD content and down-regulation of katG expression contribute to the higher resistance to INH in mc(2)155-MS5175. | ||||||||||||
| Key Molecule: D-inositol 3-phosphate glycosyltransferase (MSHA) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Non-synonymous mutation | p.F355S |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Key Molecule: D-inositol 3-phosphate glycosyltransferase (MSHA) | [10] | ||||||||||||
| Resistant Disease | Mycolicibacterium smegmatis infection [ICD-11: 1B2Z.6] | ||||||||||||
| Molecule Alteration | Non-synonymous mutation | p.N111S |
|||||||||||
| Experimental Note | Discovered Using In-vivo Testing Model | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis strain H37Rv ATCC27294 T | 83332 | |||||||||||
| Experiment for Molecule Alteration |
Sequencing analysis | ||||||||||||
| Experiment for Drug Resistance |
In vitro drug susceptibility testing | ||||||||||||
| Mechanism Description | Notably, isoniazid is activated by the enzyme catalase-peroxidase, KatG, encoded by katG, whereas prothionamide is activated by the flavin monoxygenase, EthA, encoded by ethA. Mutations in katG and ethA are associated with individual isoniazid and prothionamide/ethionamide resistance, respectively. The ndh gene coding for NADH dehydrogenase, Ndh, was first identified as a new mechanism for INHR in Mycobacterium smegmatis. The mutations in ndh gene cause defects in the oxidation of NADH to NAD, which results in NADH accumulation and NAD depletion. The increased level of NADH inhibits the binding of isoniazid-NAD adduct to the active site of the InhA enzyme, which disturbs the regulation of enzyme activity and may cause co-resistance to isoniazid and prothionamide. EthR, a member of the TetR/CamR family, is a repressor of ethA. EthR regulates the transcription of ethA by coordinated octamerization on a 55-bp operator situated in the ethA-R intergenic region. Impeding EthR function leads to enhanced mycobacterial sensitivity to prothionamide, whereas mutations in ethR encoding a negative transcriptional regulator of the expression of EthA lead to prothionamide resistance. Finally, MshA, a member of the glycosyltransferase family, is a key enzyme involved in mycothiol biosynthesis in M. tuberculosis. Mutations in mshA coding MshA have been proposed to create a disturbance in prothionamide/ethionamide activation. | ||||||||||||
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Catalase-peroxidase (KATG) | [11] | |||
| Resistant Disease | HIV-infected patients with tuberculosis [ICD-11: 1C60.0] | |||
| Molecule Alteration | Expression | Down-regulation |
||
| Experimental Note | Discovered Using In-vivo Testing Model | |||
| Cell Pathway Regulation | Cell growth | Inhibition | hsa05200 | |
| In Vitro Model | Mycobacterium smegmatis mc2155 | 246196 | ||
| Mycobacterium smegmatis mc2155-Cu | 246196 | |||
| Experiment for Molecule Alteration |
qPCR | |||
| Experiment for Drug Resistance |
MIC assay | |||
| Mechanism Description | As a prodrug, INH needs to be activated by katG to execute its antibiotic function. katG is a bifunctional enzyme with both catalase and peroxidase activity and catalyzes the coupling of INH with NAD+ to form the isonicotinic acyl-NAD complex, which binds to the enoyl-acyl carrier protein reductase to inhibit the synthesis of mycolic acid required for the mycobacterial cell wall. In the present study, quantitative proteomic analysis showed that the expression level of katG was down-regulated in mc2155-Cu as compared to mc2155. Down-regulation of katG expression as well as a decrease in cellular NAD level results in the higher resistance to INH in mc2155-Cu. | |||
| Key Molecule: Arylamine N-acetyltransferase 1 (NAT1) | [12] | |||
| Resistant Disease | HIV-infected patients with tuberculosis [ICD-11: 1C60.0] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Discovered Using In-vivo Testing Model | |||
| Cell Pathway Regulation | Cell growth | Activation | hsa05200 | |
| In Vitro Model | Mycobacterium tuberculosis H37Rv | 83332 | ||
| Experiment for Molecule Alteration |
SDS-PAGE assay | |||
| Experiment for Drug Resistance |
Titertek multiskan assay | |||
| Mechanism Description | Arylamine N-acetyltransferase (NAT), a drug-metabolizing enzyme of MTB, can acetylate INH, transferring an acetyl group from acetyl coenzyme A to the terminal nitrogen of the drug, which in its N-acetylated form is therapeutically inactive. The overexpression of NAT in Mycobacterium smegmatis showed increased resistance to INH; in addition, when the gene was knocked-out, the bacteria exhibited increased sensitivity to INH. | |||
|
|
||||
| Key Molecule: Isocitrate lyase 1 (ICL1) | [9] | |||
| Resistant Disease | HIV-infected patients with tuberculosis [ICD-11: 1C60.0] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Discovered Using In-vivo Testing Model | |||
| In Vitro Model | Mycobacterium tuberculosis strains | 1773 | ||
| Experiment for Molecule Alteration |
qRT-PCR | |||
| Experiment for Drug Resistance |
MIC assay | |||
| Mechanism Description | Despite targeting diverse cellular processes, all three drugs trigger activation of Mtb's isocitrate lyases (ICLs), metabolic enzymes commonly assumed to be involved in replenishing of tricarboxylic acid (TCA) cycle intermediates. We further show that ICL-deficient Mtb strains are significantly more susceptible than wild-type Mtb to all three antibiotics, and that this susceptibility can be chemically rescued when Mtb is co-incubated with an antioxidant. | |||
| Key Molecule: Isocitrate lyase 2 (ICL2) | [9] | |||
| Resistant Disease | HIV-infected patients with tuberculosis [ICD-11: 1C60.0] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Discovered Using In-vivo Testing Model | |||
| In Vitro Model | Mycobacterium tuberculosis strains | 1773 | ||
| Experiment for Molecule Alteration |
qRT-PCR | |||
| Experiment for Drug Resistance |
MIC assay | |||
| Mechanism Description | Despite targeting diverse cellular processes, all three drugs trigger activation of Mtb's isocitrate lyases (ICLs), metabolic enzymes commonly assumed to be involved in replenishing of tricarboxylic acid (TCA) cycle intermediates. We further show that ICL-deficient Mtb strains are significantly more susceptible than wild-type Mtb to all three antibiotics, and that this susceptibility can be chemically rescued when Mtb is co-incubated with an antioxidant. | |||
| Drug Sensitivity Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Isocitrate lyase 1 (ICL1) | [9] | |||
| Sensitive Disease | HIV-infected patients with tuberculosis [ICD-11: 1C60.0] | |||
| Molecule Alteration | Expression | Down-regulation |
||
| Experimental Note | Discovered Using In-vivo Testing Model | |||
| In Vitro Model | Mycobacterium tuberculosis strains | 1773 | ||
| Experiment for Molecule Alteration |
qRT-PCR | |||
| Experiment for Drug Resistance |
MIC assay | |||
| Mechanism Description | Despite targeting diverse cellular processes, all three drugs trigger activation of Mtb's isocitrate lyases (ICLs), metabolic enzymes commonly assumed to be involved in replenishing of tricarboxylic acid (TCA) cycle intermediates. We further show that ICL-deficient Mtb strains are significantly more susceptible than wild-type Mtb to all three antibiotics, and that this susceptibility can be chemically rescued when Mtb is co-incubated with an antioxidant. | |||
| Key Molecule: Isocitrate lyase 2 (ICL2) | [9] | |||
| Sensitive Disease | HIV-infected patients with tuberculosis [ICD-11: 1C60.0] | |||
| Molecule Alteration | Expression | Down-regulation |
||
| Experimental Note | Discovered Using In-vivo Testing Model | |||
| In Vitro Model | Mycobacterium tuberculosis strains | 1773 | ||
| Experiment for Molecule Alteration |
qRT-PCR | |||
| Experiment for Drug Resistance |
MIC assay | |||
| Mechanism Description | Despite targeting diverse cellular processes, all three drugs trigger activation of Mtb's isocitrate lyases (ICLs), metabolic enzymes commonly assumed to be involved in replenishing of tricarboxylic acid (TCA) cycle intermediates. We further show that ICL-deficient Mtb strains are significantly more susceptible than wild-type Mtb to all three antibiotics, and that this susceptibility can be chemically rescued when Mtb is co-incubated with an antioxidant. | |||
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Catalase-peroxidase (KATG) | [8] | |||
| Resistant Disease | Urinary tuberculosis [ICD-11: 1G80.0] | |||
| Molecule Alteration | Missense mutation | p.S531L |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Mycobacterium tuberculosis isolates | 1773 | ||
| Experiment for Molecule Alteration |
Gene sequencing assay | |||
| Mechanism Description | Regarding drug-resistance mutation profiles, the most prevalent mutation sites were katG S315T1 and rpoB S531L. | |||
| Key Molecule: DNA-directed RNA polymerase subunit beta (RPOB) | [8] | |||
| Resistant Disease | Urinary tuberculosis [ICD-11: 1G80.0] | |||
| Molecule Alteration | Missense mutation | p.S531L |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Mycobacterium tuberculosis isolates | 1773 | ||
| Experiment for Molecule Alteration |
Gene sequencing assay | |||
| Mechanism Description | Regarding drug-resistance mutation profiles, the most prevalent mutation sites were katG S315T1 and rpoB S531L. | |||
| Key Molecule: Catalase-peroxidase (KATG) | [8] | |||
| Resistant Disease | Urinary tuberculosis [ICD-11: 1G80.0] | |||
| Molecule Alteration | Missense mutation | p.S315T1 |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Mycobacterium tuberculosis isolates | 1773 | ||
| Experiment for Molecule Alteration |
Gene sequencing assay | |||
| Mechanism Description | Regarding drug-resistance mutation profiles, the most prevalent mutation sites were katG S315T1 and rpoB S531L. | |||
| Key Molecule: DNA-directed RNA polymerase subunit beta (RPOB) | [8] | |||
| Resistant Disease | Urinary tuberculosis [ICD-11: 1G80.0] | |||
| Molecule Alteration | Missense mutation | p.S315T1 |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Mycobacterium tuberculosis isolates | 1773 | ||
| Experiment for Molecule Alteration |
Gene sequencing assay | |||
| Mechanism Description | Regarding drug-resistance mutation profiles, the most prevalent mutation sites were katG S315T1 and rpoB S531L. | |||
ICD-12: Respiratory system diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | |||||||||||||
|
|
|||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [4] | ||||||||||||
| Resistant Disease | Pneumoconiosis complicated with tuberculosis [ICD-11: CA60.Y] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S315T |
|||||||||||
| Wild Type Structure | Method: X-ray diffraction | Resolution: 2.00 Å | |||||||||||
| Mutant Type Structure | Method: X-ray diffraction | Resolution: 2.10 Å | |||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
M
M
P
P
E
E
Q
Q
H
H
P
P
P
P
I
I
T
T
10
|
E
E
T
T
T
T
T
T
G
G
A
A
A
A
S
S
N
N
G
G
20
|
C
C
P
P
V
V
V
V
G
G
H
H
M
M
K
K
Y
Y
P
P
30
|
V
V
E
E
G
G
G
G
G
G
N
N
Q
Q
D
D
W
W
W
W
40
|
P
P
N
N
R
R
L
L
N
N
L
L
K
K
V
V
L
L
H
H
50
|
Q
Q
N
N
P
P
A
A
V
V
A
A
D
D
P
P
M
M
G
G
60
|
A
A
A
A
F
F
D
D
Y
Y
A
A
A
A
E
E
V
V
A
A
70
|
T
T
I
I
D
D
V
V
D
D
A
A
L
L
T
T
R
R
D
D
80
|
I
I
E
E
E
E
V
V
M
M
T
T
T
T
S
S
Q
Q
P
P
90
|
W
W
W
W
P
P
A
A
D
D
Y
Y
G
G
H
H
Y
Y
G
G
100
|
P
P
L
L
F
F
I
I
R
R
M
M
A
A
W
W
H
H
A
A
110
|
A
A
G
G
T
T
Y
Y
R
R
I
I
H
H
D
D
G
G
R
R
120
|
G
G
G
G
A
A
G
G
G
G
G
G
M
M
Q
Q
R
R
F
F
130
|
A
A
P
P
L
L
N
N
S
S
W
W
P
P
D
D
N
N
A
A
140
|
S
S
L
L
D
D
K
K
A
A
R
R
R
R
L
L
L
L
W
W
150
|
P
P
V
V
K
K
K
K
K
K
Y
Y
G
G
K
K
K
K
L
L
160
|
S
S
W
W
A
A
D
D
L
L
I
I
V
V
F
F
A
A
G
G
170
|
N
N
C
C
A
A
L
L
E
E
S
S
M
M
G
G
F
F
K
K
180
|
T
T
F
F
G
G
F
F
G
G
F
F
G
G
R
R
V
V
D
D
190
|
Q
Q
W
W
E
E
P
P
D
D
E
E
V
V
Y
Y
W
W
G
G
200
|
K
K
E
E
A
A
T
T
W
W
L
L
G
G
D
D
E
E
R
R
210
|
Y
Y
S
S
G
G
K
K
R
R
D
D
L
L
E
E
N
N
P
P
220
|
L
L
A
A
A
A
V
V
Q
Q
M
M
G
G
L
L
I
I
Y
Y
230
|
V
V
N
N
P
P
E
E
G
G
P
P
N
N
G
G
N
N
P
P
240
|
D
D
P
P
M
M
A
A
A
A
A
A
V
V
D
D
I
I
R
R
250
|
E
E
T
T
F
F
R
R
R
R
M
M
A
A
M
M
N
N
D
D
260
|
V
V
E
E
T
T
A
A
A
A
L
L
I
I
V
V
G
G
G
G
270
|
H
H
T
T
F
F
G
G
K
K
T
T
H
H
G
G
A
A
G
G
280
|
P
P
A
A
D
D
L
L
V
V
G
G
P
P
E
E
P
P
E
E
290
|
A
A
A
A
P
P
L
L
E
E
Q
Q
M
M
G
G
L
L
G
G
300
|
W
W
K
K
S
S
S
S
Y
Y
G
G
T
T
G
G
T
T
G
G
310
|
K
K
D
D
A
A
I
I
T
T
S
T
G
G
I
I
E
E
V
V
320
|
V
V
W
W
T
T
N
N
T
T
P
P
T
T
K
K
W
W
D
D
330
|
N
N
S
S
F
F
L
L
E
E
I
I
L
L
Y
Y
G
G
Y
Y
340
|
E
E
W
W
E
E
L
L
T
T
K
K
S
S
P
P
A
A
G
G
350
|
A
A
W
W
Q
Q
Y
Y
T
T
A
A
K
K
D
D
G
G
A
A
360
|
G
G
A
A
G
G
T
T
I
I
P
P
D
D
P
P
F
F
G
G
370
|
G
G
P
P
G
G
R
R
S
S
P
P
T
T
M
M
L
L
A
A
380
|
T
T
D
D
L
L
S
S
L
L
R
R
V
V
D
D
P
P
I
I
390
|
Y
Y
E
E
R
R
I
I
T
T
R
R
R
R
W
W
L
L
E
E
400
|
H
H
P
P
E
E
E
E
L
L
A
A
D
D
E
E
F
F
A
A
410
|
K
K
A
A
W
W
Y
Y
K
K
L
L
I
I
H
H
R
R
D
D
420
|
M
M
G
G
P
P
V
V
A
A
R
R
Y
Y
L
L
G
G
P
P
430
|
L
L
V
V
P
P
K
K
Q
Q
T
T
L
L
L
L
W
W
Q
Q
440
|
D
D
P
P
V
V
P
P
A
A
V
V
S
S
H
H
D
D
L
L
450
|
V
V
G
G
E
E
A
A
E
E
I
I
A
A
S
S
L
L
K
K
460
|
S
S
Q
Q
I
I
R
R
A
A
S
S
G
G
L
L
T
T
V
V
470
|
S
S
Q
Q
L
L
V
V
S
S
T
T
A
A
W
W
A
A
A
A
480
|
A
A
S
S
S
S
F
F
R
R
G
G
S
S
D
D
K
K
R
R
490
|
G
G
G
G
A
A
N
N
G
G
G
G
R
R
I
I
R
R
L
L
500
|
Q
Q
P
P
Q
Q
V
V
G
G
W
W
E
E
V
V
N
N
D
D
510
|
P
P
D
D
G
G
D
D
L
L
R
R
K
K
V
V
I
I
R
R
520
|
T
T
L
L
E
E
E
E
I
I
Q
Q
E
E
S
S
F
F
N
N
530
|
S
S
A
A
A
A
P
P
G
G
N
N
I
I
K
K
V
V
S
S
540
|
F
F
A
A
D
D
L
L
V
V
V
V
L
L
G
G
G
G
C
C
550
|
A
A
A
A
I
I
E
E
K
K
A
A
A
A
K
K
A
A
A
A
560
|
G
G
H
H
N
N
I
I
T
T
V
V
P
P
F
F
T
T
P
P
570
|
G
G
R
R
T
T
D
D
A
A
S
S
Q
Q
E
E
Q
Q
T
T
580
|
D
D
V
V
E
E
S
S
F
F
A
A
V
V
L
L
E
E
P
P
590
|
K
K
A
A
D
D
G
G
F
F
R
R
N
N
Y
Y
L
L
G
G
600
|
K
K
G
G
N
N
P
P
L
L
P
P
A
A
E
E
Y
Y
M
M
610
|
L
L
L
L
D
D
K
K
A
A
N
N
L
L
L
L
T
T
L
L
620
|
S
S
A
A
P
P
E
E
M
M
T
T
V
V
L
L
V
V
G
G
630
|
G
G
L
L
R
R
V
V
L
L
G
G
A
A
N
N
Y
Y
K
K
640
|
R
R
L
L
P
P
L
L
G
G
V
V
F
F
T
T
E
E
A
A
650
|
S
S
E
E
S
S
L
L
T
T
N
N
D
D
F
F
F
F
V
V
660
|
N
N
L
L
L
L
D
D
M
M
G
G
I
I
T
T
W
W
E
E
670
|
P
P
S
S
P
P
A
A
D
D
D
D
G
G
T
T
Y
Y
Q
Q
680
|
G
G
K
K
D
D
G
G
S
S
G
G
K
K
V
V
K
K
W
W
690
|
T
T
G
G
S
S
R
R
V
V
D
D
L
L
V
V
F
F
G
G
700
|
S
S
N
N
S
S
E
E
L
L
R
R
A
A
L
L
V
V
E
E
710
|
V
V
Y
Y
G
G
A
A
D
D
D
D
A
A
Q
Q
P
P
K
K
720
|
F
F
V
V
Q
Q
D
D
F
F
V
V
A
A
A
A
W
W
D
D
730
|
K
K
V
V
M
M
N
N
L
L
D
D
R
R
F
F
D
D
V
V
740
|
R
R
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis isolates | 1773 | |||||||||||
| Mycobacterium tuberculosis H37Rv1 | 1773 | ||||||||||||
| Experiment for Molecule Alteration |
qRT-PCR | ||||||||||||
| Mechanism Description | Isoniazid is a hydrazine chemical synthetic drug, which is able to be oxidized to isonicotinic acid by the catalase-peroxidase encoded by the katG gene that participates in the synthesis of coenzyme I (NAD) to inhibit the biosynthesis of mycolic acid of the cell wall in Mycobacterium tuberculosis, so as to damage the MDR-TB's barricade of resisting antioxygen and invasion. Due to deletion or mutation in the katG gene, resistance is able to be generated as the enzymatic activity is lost or degraded, thus, inhibiting the activation of Isoniazid. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [4] | ||||||||||||
| Resistant Disease | Pneumoconiosis complicated with tuberculosis [ICD-11: CA60.Y] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S315N |
|||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis isolates | 1773 | |||||||||||
| Mycobacterium tuberculosis H37Rv1 | 1773 | ||||||||||||
| Experiment for Molecule Alteration |
qRT-PCR | ||||||||||||
| Mechanism Description | Isoniazid is a hydrazine chemical synthetic drug, which is able to be oxidized to isonicotinic acid by the catalase-peroxidase encoded by the katG gene that participates in the synthesis of coenzyme I (NAD) to inhibit the biosynthesis of mycolic acid of the cell wall in Mycobacterium tuberculosis, so as to damage the MDR-TB's barricade of resisting antioxygen and invasion. Due to deletion or mutation in the katG gene, resistance is able to be generated as the enzymatic activity is lost or degraded, thus, inhibiting the activation of Isoniazid. | ||||||||||||
| Key Molecule: Catalase-peroxidase (KATG) | [4] | ||||||||||||
| Resistant Disease | Pneumoconiosis complicated with tuberculosis [ICD-11: CA60.Y] | ||||||||||||
| Molecule Alteration | Missense mutation | p.A431V |
|||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | Mycobacterium tuberculosis isolates | 1773 | |||||||||||
| Mycobacterium tuberculosis H37Rv1 | 1773 | ||||||||||||
| Experiment for Molecule Alteration |
qRT-PCR | ||||||||||||
| Mechanism Description | Isoniazid is a hydrazine chemical synthetic drug, which is able to be oxidized to isonicotinic acid by the catalase-peroxidase encoded by the katG gene that participates in the synthesis of coenzyme I (NAD) to inhibit the biosynthesis of mycolic acid of the cell wall in Mycobacterium tuberculosis, so as to damage the MDR-TB's barricade of resisting antioxygen and invasion. Due to deletion or mutation in the katG gene, resistance is able to be generated as the enzymatic activity is lost or degraded, thus, inhibiting the activation of Isoniazid. | ||||||||||||
ICD-X: Extension Codes
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Catalase (CAT) | [2] | |||
| Resistant Disease | bifidobacterium adolescentis infection [ICD-11: XN33F] | |||
| Molecule Alteration | Function | RV0005; p.Ala403Ser |
||
| Experimental Note | Revealed Based on the Cell Line Data | |||
| In Vitro Model | Bifidobacterial strains | 1763 | ||
| Experiment for Molecule Alteration |
PCR; Catalase foam assay; Catalase gel assay | |||
| Experiment for Drug Resistance |
Growth curve assay; Spot assay; Anti-tubercular drug uptake and surface assay; Adaptability assay; FE-SEM assay; MIC assay; Particle size assay | |||
| Mechanism Description | The current study aims to understand the resistance of Bifidobacterium adolescentis to different anti-tubercular drugs (first-line oral tuberculosis drugs). The bacteria were grown with anti-tubercular drugs such as isoniazid, pyrazinamide, and streptomycin to better understand the resistance phenomena. It was found that even at tenfold higher concentrations, growth rates remained unchanged. In addition, a small number of bacteria were found to aggregate strongly, a property that protects against the toxicity of the drug. Further FE-SEM (Field Emission Scanning Electron Microscopy) analysis revealed that some bacteria became excessively long, elongated, and protruded on the surface. Size scattering analysis confirmed the presence of bifidobacteria in the size range of 1.0-100 um. After whole genome sequence analysis, certain mutations were found in the relevant gene. In vitro, foam formation and growth in the presence of H2O2 and HPLC (High Performance Liquid Chromatography) studies provide additional evidence for the presence of catalase. According to RAST (Rapid Annotation Using Subsystems Technology) annotation and CARD (Comprehensive Antibiotic Resistance Database analysis), there were not many components in the genome that were resistant to antibiotics. Whole genome sequence (WGS) analysis does not show the presence of bacteriocins and antibiotic resistance genes, but few hypothetical proteins were observed. 3D structure and docking studies suggest their interaction with specific ligands. | |||
References
If you find any error in data or bug in web service, please kindly report it to Dr. Sun and Dr. Yu.
