Drug Information
Drug (ID: DG00005) and It's Reported Resistant Information
| Name |
Benzylpenicillin
|
||||
|---|---|---|---|---|---|
| Synonyms |
Penicillin g; Benzylpenicillinic acid; Free penicillin II; Pencillin G; 61-33-6; Benzylpenicillin G; Bencilpenicilina; Benzylpenicillinum; Benzyl penicillin; Free penicillin G; Dropcillin; Pharmacillin; Gelacillin; Liquacillin; Benzopenicillin; Cilopen; Pradupen; Benzylpenicilline; Specilline G; Cilloral; Cosmopen; Ursopen; Galofak; Free benzylpenicillin; Compocillin G; Phenylacetamidopenicillanic acid; 6-(2-Phenylacetamido)penicillanic acid; Pfizerpen; Benzyl-6-aminopenicillinic acid
Click to Show/Hide
|
||||
| Structure |
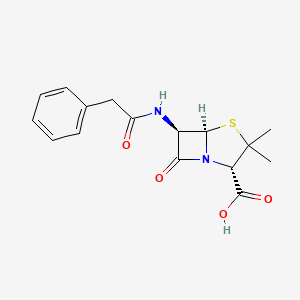
|
||||
| Drug Resistance Disease(s) |
Disease(s) with Resistance Information Discovered by Cell Line Test for This Drug
(1 diseases)
[1]
Disease(s) with Clinically Reported Resistance for This Drug
(3 diseases)
[4]
[5]
Disease(s) with Resistance Information Validated by in-vivo Model for This Drug
(1 diseases)
[6]
|
||||
| Target | Bacterial Penicillin binding protein 1 (Bact pbp) |
PBPA_ECOLI
; PBPB_ECOLI ; PBPC_ECOLI |
[2] | ||
| Bacterial Penicillin binding protein 3 (Bact mrcA) | FTSI_ECOLI | [2] | |||
| Click to Show/Hide the Molecular Information and External Link(s) of This Drug | |||||
| Formula |
C16H18N2O4S
|
||||
| IsoSMILES |
CC1([C@@H](N2[C@H](S1)[C@@H](C2=O)NC(=O)CC3=CC=CC=C3)C(=O)O)C
|
||||
| InChI |
1S/C16H18N2O4S/c1-16(2)12(15(21)22)18-13(20)11(14(18)23-16)17-10(19)8-9-6-4-3-5-7-9/h3-7,11-12,14H,8H2,1-2H3,(H,17,19)(H,21,22)/t11-,12+,14-/m1/s1
|
||||
| InChIKey |
JGSARLDLIJGVTE-MBNYWOFBSA-N
|
||||
| PubChem CID | |||||
| ChEBI ID | |||||
| TTD Drug ID | |||||
| VARIDT ID | |||||
| INTEDE ID | |||||
| DrugBank ID | |||||
Type(s) of Resistant Mechanism of This Drug
Drug Resistance Data Categorized by Their Corresponding Diseases
ICD-01: Infectious/parasitic diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Beta-lactamase (BLA) | [6] | |||
| Resistant Disease | Rhodobacter sphaeroides infection [ICD-11: 1A00-1C4Z] | |||
| Molecule Alteration | Expression | Inherence |
||
| Experimental Note | Discovered Using In-vivo Testing Model | |||
| In Vitro Model | Rhodopseudomonas sphaeroides strain DSM 160(Y) | 1063 | ||
| Rhodopseudomonas sphaeroides strain DSM158 | 1063 | |||
| Rhodopseudomonas sphaeroides strain DSM159 | 1063 | |||
| Experiment for Molecule Alteration |
Sodium dodecyl sulfate-PAGE assay | |||
| Experiment for Drug Resistance |
MIC assay | |||
| Mechanism Description | Thirteen strains of the gram-negative, facultative phototrophic bacterium Rhodobacter sphaeroides were examined fro susceptibility to beta-lactam antibiotics. All strains were sensitive to the semisynthetic penicillins ampicillin, carbenicillin, oxacillin, cloxacillin, and methicillin, but 10 of the 13 strains were resistant to penicillin G, as well as a number of cephalosporins, such as cephalothin, cephapirin, and cephalosporin C. A beta-lactamase (EC 3.5.2.6) with strong cephalosporinase activity was detected in all of the resistant strains of R. sphaeroides. With strain Y-1 as a model, it was shown that the beta-lactamase was inducible by penicillin G, cephalosporin C, cephalothin, and to some minor extent, cephapirin. | |||
| Key Molecule: KBL-1 protein (KBL-1) | [1] | |||
| Resistant Disease | Lactobacillus casei infection [ICD-11: 1A00-1C4Z] | |||
| Molecule Alteration | Expression | . |
||
| Experimental Note | Revealed Based on the Cell Line Data | |||
| In Vitro Model | S. maltophilia JUNP497 | N.A. | ||
| Mechanism Description | Recombinant KBL-1 protein had hydrolytic activities against all the beta-lactams tested, except for aztreonam (Table?3). Recombinant KBL-1 efficiently hydrolyzed the penicillins, including ampicillin, amoxicillin, penicillin G, and piperacillin with?kcat/km?values of 0.422 to 1.166. | |||
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Outer membrane porin (OMP38) | [2], [3] | |||
| Resistant Disease | Melioidosis [ICD-11: 1C42.0] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli | 668369 | ||
| Escherichia coli BL21(DE3) | 469008 | |||
| Burkholderia pseudomallei isolates | 28450 | |||
| Experiment for Molecule Alteration |
Whole genome sequence assay | |||
| Experiment for Drug Resistance |
Broth microdilution method assay | |||
| Mechanism Description | Bps is highly resistant to many antimicrobial agents and this resistance may result from the low drug permeability of outer membrane proteins, known as porins.An Escherichia coli strain defective in most porins, but expressing BpsOmp38, exhibited considerably lower antimicrobial susceptibility than the control strain. In addition, mutation of Tyr119, the most prominent pore-lining residue in BpsOmp38, markedly altered membrane permeability, substitution with Ala (mutant BpsOmp38Y119A) enhanced uptake of the antimicrobial agents, while substitution with Phe (mutant BpsOmp38Y119F) inhibited uptake. | |||
ICD-12: Respiratory system diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Bcr/CflA family efflux transporter (BCML) | [4] | |||
| Resistant Disease | Klebsiella pneumoniae infection [ICD-11: CA40.1] | |||
| Molecule Alteration | Expression | Inherence |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli DH10B | 316385 | ||
| Escherichia coli strain NCTC 50192 | 562 | |||
| Klebsiella pneumoniae strain ORI-1 | 573 | |||
| Experiment for Molecule Alteration |
PCR and hybridization experiments assay | |||
| Experiment for Drug Resistance |
Agar dilution technique assay | |||
| Mechanism Description | Klebsiella pneumoniae ORI-1 strain harbored a ca. 140-kb nontransferable plasmid, pTk1, that conferred an extended-spectrum cephalosporin resistance profile antagonized by the addition of clavulanic acid, tazobactam, or imipenem. The gene for GES-1 (Guiana extended-spectrum beta-lactamase) was cloned, and its protein was expressed in Escherichia coli DH10B, where this pI-5. 8 beta-lactamase of a ca. 31-kDa molecular mass conferred resistance to oxyimino cephalosporins (mostly to ceftazidime). GES-1 is weakly related to the other plasmid-located Ambler class A extended-spectrum beta-lactamases (ESBLs). | |||
| Key Molecule: Bcr/CflA family efflux transporter (BCML) | [4] | |||
| Resistant Disease | Klebsiella pneumoniae infection [ICD-11: CA40.1] | |||
| Molecule Alteration | Expression | Acquired |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli DH10B | 316385 | ||
| Escherichia coli strain NCTC 50192 | 562 | |||
| Klebsiella pneumoniae strain ORI-1 | 573 | |||
| Experiment for Molecule Alteration |
PCR and hybridization experiments assay | |||
| Experiment for Drug Resistance |
Agar dilution technique assay | |||
| Mechanism Description | Beta-Lactam MICs for k. pneumoniae ORI-1 and Escherichia coli DH10B harboring either the natural plasmid pTk1 or the recombinant plasmid pC1 were somewhat similar and might indicate the presence of an ESBL. In all cases, the ceftazidime MICs were higher than those of cefotaxime and aztreonam. Beta-Lactam MICs were always lowered by the addition of clavulanic acid or tazobactam, less so by sulbactam, and uncommonly by imipenem. | |||
| Drug Sensitivity Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Bcr/CflA family efflux transporter (BCML) | [4] | |||
| Sensitive Disease | Klebsiella pneumoniae infection [ICD-11: CA40.1] | |||
| Molecule Alteration | Expression | Antagonism |
||
| Experimental Note | Identified from the Human Clinical Data | |||
| In Vitro Model | Escherichia coli DH10B | 316385 | ||
| Escherichia coli strain NCTC 50192 | 562 | |||
| Klebsiella pneumoniae strain ORI-1 | 573 | |||
| Experiment for Molecule Alteration |
PCR and hybridization experiments assay | |||
| Experiment for Drug Resistance |
Agar dilution technique assay | |||
| Mechanism Description | Inhibition studies, as measured by IC50 values with benzylpenicillin as the substrate, showed that GES-1 was inhibited by clavulanic acid (5 uM) and tazobactam (2.5 uM) and strongly inhibited by imipenem (0.1 uM). Beta-Lactam MICs were always lowered by the addition of clavulanic acid or tazobactam, less so by sulbactam, and uncommonly by imipenem. | |||
References
If you find any error in data or bug in web service, please kindly report it to Dr. Sun and Dr. Yu.
