Drug Information
Drug (ID: DG02028) and It's Reported Resistant Information
| Name |
Dichloroacetic acid
|
||||
|---|---|---|---|---|---|
| Synonyms |
DICHLOROACETIC ACID|79-43-6|2,2-Dichloroacetic acid|Dichloracetic acid|Acetic acid, dichloro-|Dichloroethanoic acid|Urner's liquid|Dichlorethanoic acid|Bichloracetic acid|Bichloroacetic Acid|DCA (acid)|DICHLORO-ACETIC ACID|Acetic acid, 2,2-dichloro-|Kyselina dichloroctova|NSC 2654|Dichloroacetic acid (IUPAC)|Kyselina dichloroctova [Czech]|CCRIS 4016|HSDB 6894|Dichloressigsaeure|UNII-9LSH52S3LQ|NSC-2654|EINECS 201-207-0|9LSH52S3LQ|DKhUK|UN1764|BRN 1098596|DTXSID2020428|CHEBI:36386|AI3-18370|DCA|MFCD00004223|DTXCID20428|DICHLOROACETIC ACID [MI]|Dichloroacetate, Potassium|DICHLOROACETIC ACID [HSDB]|DICHLOROACETIC ACID [IARC]|DICHLOROACETIC ACID [VANDF]|4-02-00-00498 (Beilstein Handbook Reference)|DICHLOROACETIC ACID [WHO-DD]|UN 1764|NCGC00091444-02|Kyselina dichloroctova (Czech)|2,2-Dichloroacetic Acid-2-13C|DICHLOROACETIC ACID (IARC)|CAS-79-43-6|SMR000059158|TF4|Acid, Bichloroacetic|Acid, Dichloroacetic|dicloroacetic acid|WLN: QVYGG|2,2-dichloro-acetic acid|SCHEMBL7777|MLS000028893|MLS001074861|MLS001424165|CHEMBL13960|NSC2654|BDBM227588|HMS2051P22|HMS2232H11|HMS3370D12|HMS3393P22|YGA47070|Dichloroacetic acid, puriss., 99%|Tox21_111133|Tox21_202363|Tox21_300419|Acetic acid, dichloro- (8CI,9CI)|BBL011449|LMFA01090070|MSK000808|STL146561|AKOS005720869|Tox21_111133_1|AT42091|CCG-101070|DB08809|NC00320|NCGC00091444-01|NCGC00091444-03|NCGC00091444-04|NCGC00167845-02|NCGC00254261-01|NCGC00259912-01|AS-14228|DB-056364|D0308|Dichloroacetic acid [UN1764] [Corrosive]|Dichloroacetic acid, biotech. grade, >=98%|NS00001485|Dichloroacetic acid, ReagentPlus(R), >=99%|EN300-20014|Q412845|DCA Deblock (0.36M dichloroacetic acid in toluene)|Dichloroacetic acid, PESTANAL(R), analytical standard|F2191-0226|Z104476414|Dichloroacetic acid 1000 microg/mL in Methyl-tert-butyl ether|5% Dichloroacetic Acid in Toluene (for AKTA oligopilottrade mark) NC-0406 emp Biotech GmbH|201-207-0
Click to Show/Hide
|
||||
| Indication |
In total 1 Indication(s)
.
|
||||
| Structure |
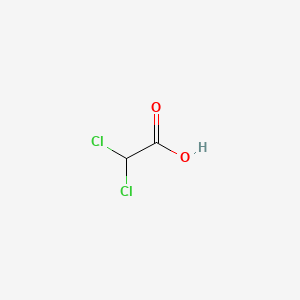
|
||||
| Click to Show/Hide the Molecular Information and External Link(s) of This Drug | |||||
| Formula |
C2H2Cl2O2
|
||||
| IsoSMILES |
C(C(=O)O)(Cl)Cl
|
||||
| InChI |
InChI=1S/C2H2Cl2O2/c3-1(4)2(5)6/h1H,(H,5,6)
|
||||
| InChIKey |
JXTHNDFMNIQAHM-UHFFFAOYSA-N
|
||||
| PubChem CID | |||||
Type(s) of Resistant Mechanism of This Drug
Drug Resistance Data Categorized by Their Corresponding Diseases
ICD-02: Benign/in-situ/malignant neoplasm
| Drug Sensitivity Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Key Molecule: Pyruvate dehydrogenase kinase 2 (PDK2) | [1] | |||
| Metabolic Type | Glucose metabolism | |||
| Sensitive Disease | Esophageal squamous cell carcinoma [ICD-11: 2B70.3] | |||
| Molecule Alteration | Expression | Up-regulation |
||
| Experimental Note | Revealed Based on the Cell Line Data | |||
| In Vivo Model | Male BALB/c nude mice (6-week old), with KYSE150 cells | Mice | ||
| Experiment for Molecule Alteration |
qRT-PCR; Western blot analysis | |||
| Experiment for Drug Resistance |
Tumor volume assay | |||
| Mechanism Description | PDK1 inhibition by siRNA or DCA significantly suppressed the growth of ESCC cells. miR-6516-5p/PDK1 axis suppressed the growth of ESCC cell by inhibiting glycolysis. Moreover, DCA and DDP synergistically inhibited the progression and glycolysis ability of ESCC cells both in vitro and in vivo by increasing oxidative stress partly through the suppression of Keap1/Nrf2 signaling pathway. | |||
References
If you find any error in data or bug in web service, please kindly report it to Dr. Sun and Dr. Yu.
