Drug Information
Drug (ID: DG00116) and It's Reported Resistant Information
| Name |
Cabotegravir
|
||||
|---|---|---|---|---|---|
| Synonyms |
Cabotegravir; 1051375-10-0; GSK1265744; UNII-HMH0132Z1Q; GSK744; GSK1265744A; GSK-1265744; HMH0132Z1Q; Cabotegravir (GSK744, GSK1265744); CAB; Oxazolo[3,2-a]pyrido[1,2-d]pyrazine-8-carboxamide, N-[(2,4-difluorophenyl)methyl]-2,3,5,7,11,11a-hexahydro-6-hydroxy-3-methyl-5,7-dioxo-, (3S,11aR)-;Oxazolo[3,2-a]pyrido[1,2-d]pyrazine-8-carboxamide, N-[(2,4-difluorophenyl)methyl]-2,3,5,7,11,11a-hexahydro-6-hydroxy-3-methyl-5,7-dioxo-, (3S,11aR)-; Cabotegravir [USAN:INN]; GSK744 LAP; GSK744 LA; S/GSK1265744; GSK 744; 744 LA; GSK 1265
Click to Show/Hide
|
||||
| Indication |
In total 1 Indication(s)
|
||||
| Structure |

|
||||
| Drug Resistance Disease(s) |
Disease(s) with Clinically Reported Resistance for This Drug
(1 diseases)
[1]
|
||||
| Target | Human immunodeficiency virus Integrase (HIV IN) | POL_HV1B1 | [1] | ||
| Click to Show/Hide the Molecular Information and External Link(s) of This Drug | |||||
| Formula |
C19H17F2N3O5
|
||||
| IsoSMILES |
C[C@H]1CO[C@H]2N1C(=O)C3=C(C(=O)C(=CN3C2)C(=O)NCC4=C(C=C(C=C4)F)F)O
|
||||
| InChI |
1S/C19H17F2N3O5/c1-9-8-29-14-7-23-6-12(16(25)17(26)15(23)19(28)24(9)14)18(27)22-5-10-2-3-11(20)4-13(10)21/h2-4,6,9,14,26H,5,7-8H2,1H3,(H,22,27)/t9-,14+/m0/s1
|
||||
| InChIKey |
WCWSTNLSLKSJPK-LKFCYVNXSA-N
|
||||
| PubChem CID | |||||
| TTD Drug ID | |||||
| INTEDE ID | |||||
| DrugBank ID | |||||
Type(s) of Resistant Mechanism of This Drug
Drug Resistance Data Categorized by Their Corresponding Diseases
ICD-01: Infectious/parasitic diseases
| Drug Resistance Data Categorized by Their Corresponding Mechanisms | |||||||||||||
|
|
|||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.E10D+p.S17T+p.S39N+p.I72V+p.S119P+p.T122I+p.T124N+p.T125A+p.V151I+p.N155H+p.N222K+p.P261A |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |
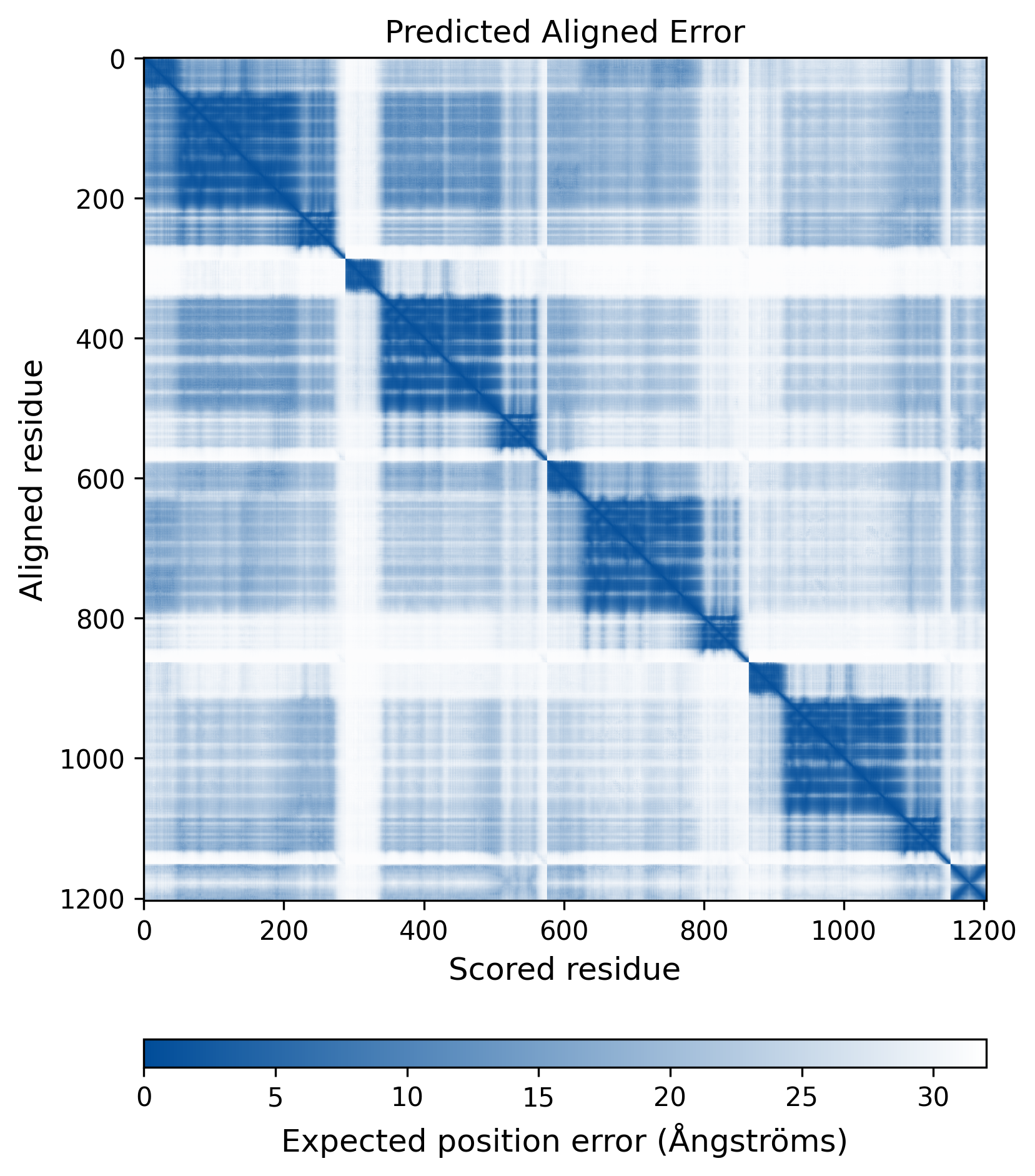
|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 74.551 |
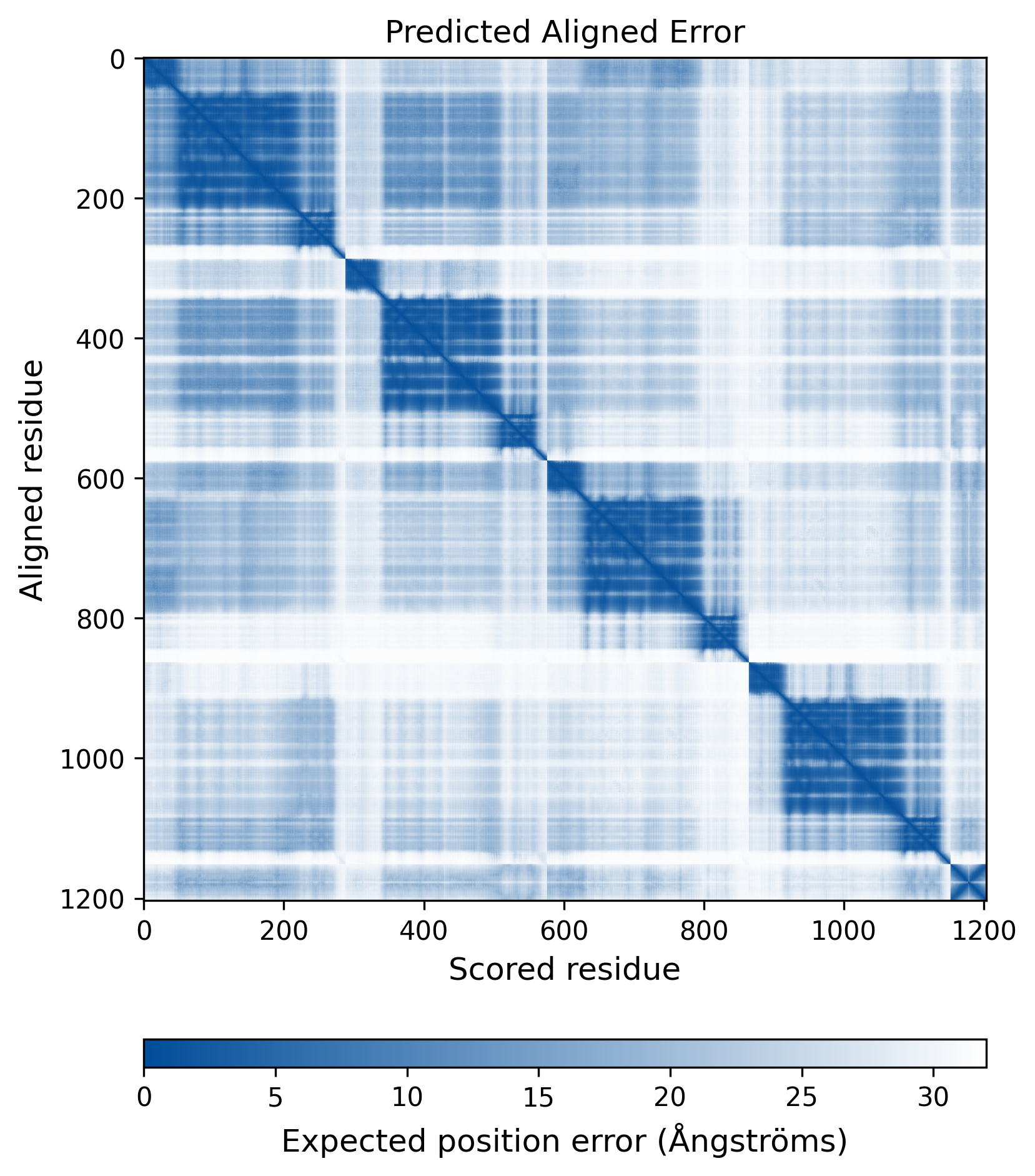
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
10
|
E
D
E
E
H
H
E
E
K
K
Y
Y
H
H
S
T
N
N
W
W
20
|
R
R
A
A
M
M
A
A
S
S
D
D
F
F
N
N
L
L
P
P
30
|
P
P
V
V
V
V
A
A
K
K
E
E
I
I
V
V
A
A
S
N
40
|
C
C
D
D
K
K
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
50
|
M
M
H
H
G
G
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
60
|
I
I
W
W
Q
Q
L
L
D
D
C
C
T
T
H
H
L
L
E
E
70
|
G
G
K
K
I
V
I
I
L
L
V
V
A
A
V
V
H
H
V
V
80
|
A
A
S
S
G
G
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
90
|
P
P
A
A
E
E
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
100
|
F
F
L
L
L
L
K
K
L
L
A
A
G
G
R
R
W
W
P
P
110
|
V
V
K
K
T
T
I
I
H
H
T
T
D
D
N
N
G
G
S
P
120
|
N
N
F
F
T
I
G
G
T
N
T
A
V
V
R
R
A
A
A
A
130
|
C
C
W
W
W
W
A
A
G
G
I
I
K
K
Q
Q
E
E
F
F
140
|
G
G
I
I
P
P
Y
Y
N
N
P
P
Q
Q
S
S
Q
Q
G
G
150
|
V
V
V
I
E
E
S
S
M
M
N
H
K
K
E
E
L
L
K
K
160
|
K
K
I
I
I
I
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
170
|
E
E
H
H
L
L
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
180
|
V
V
F
F
I
I
H
H
N
N
F
F
K
K
R
R
K
K
G
G
190
|
G
G
I
I
G
G
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
200
|
I
I
V
V
D
D
I
I
I
I
A
A
T
T
D
D
I
I
Q
Q
210
|
T
T
K
K
E
E
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
220
|
I
I
Q
Q
N
K
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
230
|
S
S
R
R
N
N
P
P
L
L
W
W
K
K
G
G
P
P
A
A
240
|
K
K
L
L
L
L
W
W
K
K
G
G
E
E
G
G
A
A
V
V
250
|
V
V
I
I
Q
Q
D
D
N
N
S
S
D
D
I
I
K
K
V
V
260
|
V
V
P
A
R
R
R
R
K
K
A
A
K
K
I
I
I
I
R
R
270
|
D
D
Y
Y
G
G
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
280
|
C
C
V
V
A
A
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q176H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.D6T+p.K7E+p.S17N+p.V54I+p.I72V+p.L74M+p.V79I+p.T97A+p.L101I+p.I113L+p.S119R+p.T122I+p.T124N+p.T125A+p.E138K+p.G140A+p.I141AT+p.S147SG+p.Q148R+p.K156N+p.I203M+p.I208M+p.L234V+p.D253E |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 73.975 |
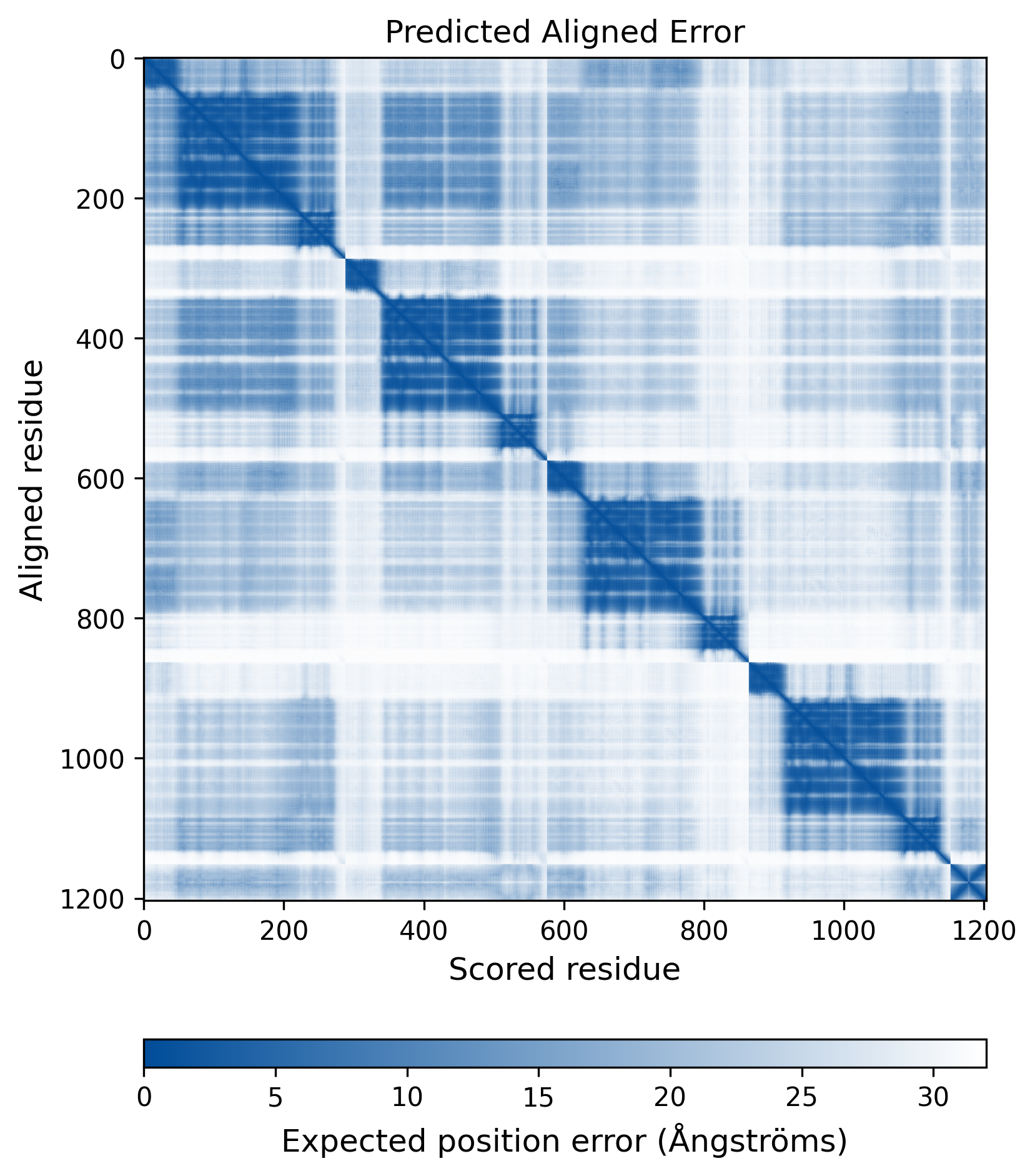
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
D
D
G
G
I
I
D
T
K
E
A
A
Q
Q
10
|
D
D
E
E
H
H
E
E
K
K
Y
Y
H
H
S
N
N
N
W
W
20
|
R
R
A
A
M
M
A
A
S
S
D
D
F
F
N
N
L
L
P
P
30
|
P
P
V
V
V
V
A
A
K
K
E
E
I
I
V
V
A
A
S
S
40
|
C
C
D
D
K
K
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
50
|
M
M
H
H
G
G
Q
Q
V
I
D
D
C
C
S
S
P
P
G
G
60
|
I
I
W
W
Q
Q
L
L
D
D
C
C
T
T
H
H
L
L
E
E
70
|
G
G
K
K
I
V
I
I
L
M
V
V
A
A
V
V
H
H
V
I
80
|
A
A
S
S
G
G
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
90
|
P
P
A
A
E
E
T
T
G
G
Q
Q
E
E
T
A
A
A
Y
Y
100
|
F
F
L
I
L
L
K
K
L
L
A
A
G
G
R
R
W
W
P
P
110
|
V
V
K
K
T
T
I
L
H
H
T
T
D
D
N
N
G
G
S
R
120
|
N
N
F
F
T
I
G
G
T
N
T
A
V
V
R
R
A
A
A
A
130
|
C
C
W
W
W
W
A
A
G
G
I
I
K
K
Q
Q
E
K
F
F
140
|
G
A
I
A
P
P
Y
Y
N
N
P
P
Q
Q
S
G
Q
R
G
G
150
|
V
V
V
V
E
E
S
S
M
M
N
N
K
N
E
E
L
L
K
K
160
|
K
K
I
I
I
I
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
170
|
E
E
H
H
L
L
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
180
|
V
V
F
F
I
I
H
H
N
N
F
F
K
K
R
R
K
K
G
G
190
|
G
G
I
I
G
G
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
200
|
I
I
V
V
D
D
I
M
I
I
A
A
T
T
D
D
I
M
Q
Q
210
|
T
T
K
K
E
E
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
220
|
I
I
Q
Q
N
N
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
230
|
S
S
R
R
N
N
P
P
L
V
W
W
K
K
G
G
P
P
A
A
240
|
K
K
L
L
L
L
W
W
K
K
G
G
E
E
G
G
A
A
V
V
250
|
V
V
I
I
Q
Q
D
E
N
N
S
S
D
D
I
I
K
K
V
V
260
|
V
V
P
P
R
R
R
R
K
K
A
A
K
K
I
I
I
I
R
R
270
|
D
D
Y
Y
G
G
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
280
|
C
C
V
V
A
A
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q184H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.K14R+p.V31I+p.V32I+p.L45Q+p.L101I+p.T112V+p.T125A+p.G134N+p.I135V+p.K136T+p.Y143C+p.V201I+p.T206S+p.S230R+p.L234I+p.N254NS+p.R269K+p.D270H |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 73.008 |
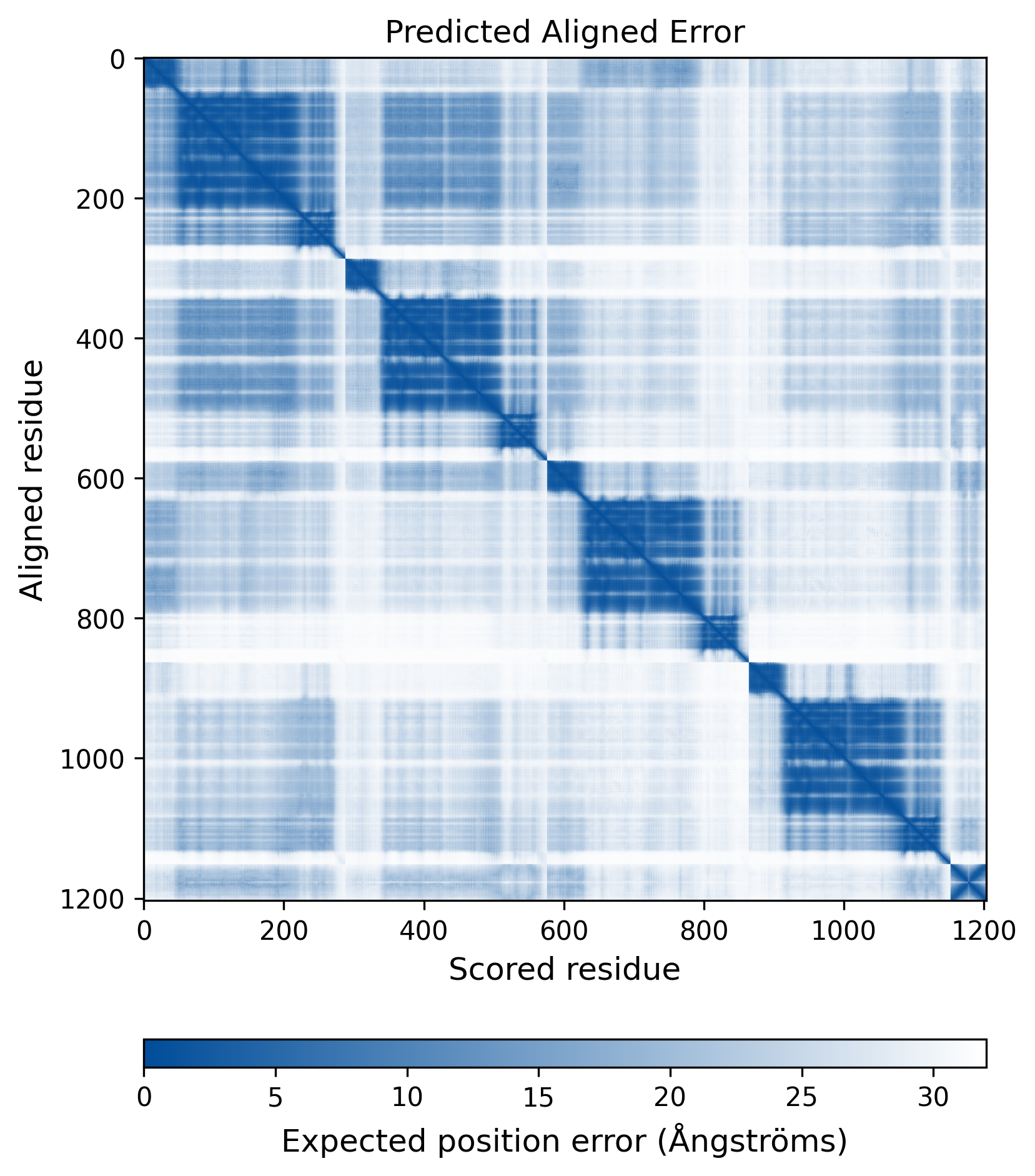
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
10
|
D
D
E
E
H
H
E
E
K
R
Y
Y
H
H
S
S
N
N
W
W
20
|
R
R
A
A
M
M
A
A
S
S
D
D
F
F
N
N
L
L
P
P
30
|
P
P
V
I
V
I
A
A
K
K
E
E
I
I
V
V
A
A
S
S
40
|
C
C
D
D
K
K
C
C
Q
Q
L
Q
K
K
G
G
E
E
A
A
50
|
M
M
H
H
G
G
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
60
|
I
I
W
W
Q
Q
L
L
D
D
C
C
T
T
H
H
L
L
E
E
70
|
G
G
K
K
V
V
I
I
L
L
V
V
A
A
V
V
H
H
V
V
80
|
A
A
S
S
G
G
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
90
|
P
P
A
A
E
E
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
100
|
F
F
L
I
L
L
K
K
L
L
A
A
G
G
R
R
W
W
P
P
110
|
V
V
K
K
T
V
I
I
H
H
T
T
D
D
N
N
G
G
S
S
120
|
N
N
F
F
T
T
G
G
A
A
T
A
V
V
R
R
A
A
A
A
130
|
C
C
W
W
W
W
A
A
G
N
I
V
K
T
Q
Q
E
E
F
F
140
|
G
G
I
I
P
P
Y
C
N
N
P
P
Q
Q
S
S
Q
Q
G
G
150
|
V
V
V
V
E
E
S
S
M
M
N
N
K
K
E
E
L
L
K
K
160
|
K
K
I
I
I
I
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
170
|
E
E
H
H
L
L
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
180
|
V
V
F
F
I
I
H
H
N
N
F
F
K
K
R
R
K
K
G
G
190
|
G
G
I
I
G
G
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
200
|
I
I
V
I
D
D
I
I
I
I
A
A
T
S
D
D
I
I
Q
Q
210
|
T
T
K
K
E
E
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
220
|
I
I
Q
Q
N
N
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
230
|
S
R
R
R
N
N
P
P
L
I
W
W
K
K
G
G
P
P
A
A
240
|
K
K
L
L
L
L
W
W
K
K
G
G
E
E
G
G
A
A
V
V
250
|
V
V
I
I
Q
Q
D
D
N
S
S
S
D
D
I
I
K
K
V
V
260
|
V
V
P
P
R
R
R
R
K
K
A
A
K
K
I
I
I
I
R
K
270
|
D
H
Y
Y
G
G
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
280
|
C
C
V
V
A
A
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q174H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.E11D+p.K14R+p.S17T+p.V31I+p.L45V+p.A91S+p.L101I+p.T124A+p.E138Q+p.G140C+p.Q148R+p.V151I+p.K219N+p.N222K+p.L234I+p.V259I |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 74.772 |
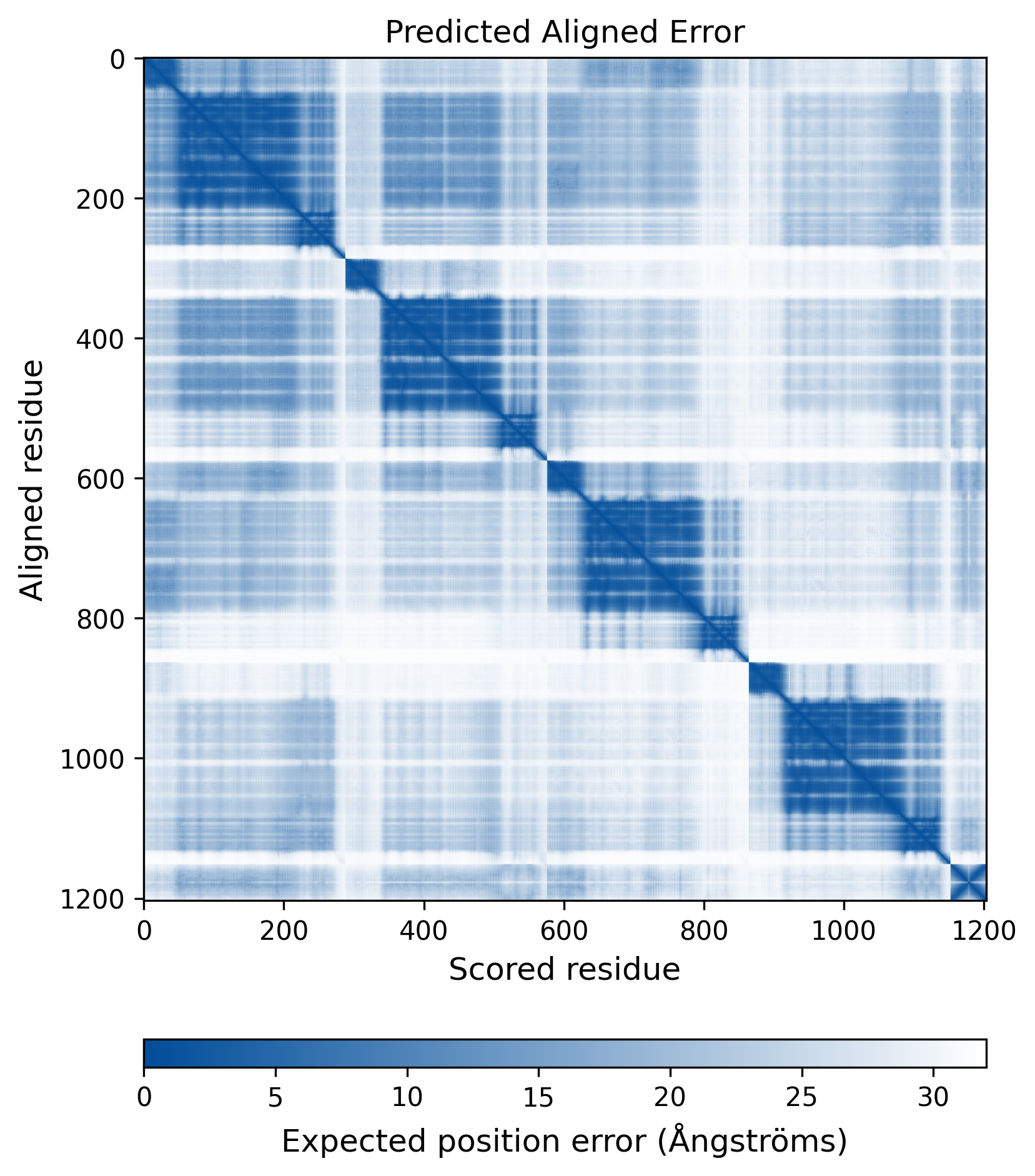
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
10
|
D
D
E
D
H
H
E
E
K
R
Y
Y
H
H
S
T
N
N
W
W
20
|
R
R
A
A
M
M
A
A
S
S
D
D
F
F
N
N
L
L
P
P
30
|
P
P
V
I
V
V
A
A
K
K
E
E
I
I
V
V
A
A
S
S
40
|
C
C
D
D
K
K
C
C
Q
Q
L
V
K
K
G
G
E
E
A
A
50
|
M
M
H
H
G
G
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
60
|
I
I
W
W
Q
Q
L
L
D
D
C
C
T
T
H
H
L
L
E
E
70
|
G
G
K
K
V
V
I
I
L
L
V
V
A
A
V
V
H
H
V
V
80
|
A
A
S
S
G
G
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
90
|
P
P
A
S
E
E
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
100
|
F
F
L
I
L
L
K
K
L
L
A
A
G
G
R
R
W
W
P
P
110
|
V
V
K
K
T
T
I
I
H
H
T
T
D
D
N
N
G
G
S
S
120
|
N
N
F
F
T
T
G
G
T
A
T
T
V
V
R
R
A
A
A
A
130
|
C
C
W
W
W
W
A
A
G
G
I
I
K
K
Q
Q
E
Q
F
F
140
|
G
C
I
I
P
P
Y
Y
N
N
P
P
Q
Q
S
S
Q
R
G
G
150
|
V
V
V
I
E
E
S
S
M
M
N
N
K
K
E
E
L
L
K
K
160
|
K
K
I
I
I
I
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
170
|
E
E
H
H
L
L
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
180
|
V
V
F
F
I
I
H
H
N
N
F
F
K
K
R
R
K
K
G
G
190
|
G
G
I
I
G
G
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
200
|
I
I
V
V
D
D
I
I
I
I
A
A
T
T
D
D
I
I
Q
Q
210
|
T
T
K
K
E
E
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
N
220
|
I
I
Q
Q
N
K
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
230
|
S
S
R
R
N
N
P
P
L
I
W
W
K
K
G
G
P
P
A
A
240
|
K
K
L
L
L
L
W
W
K
K
G
G
E
E
G
G
A
A
V
V
250
|
V
V
I
I
Q
Q
D
D
N
N
S
S
D
D
I
I
K
K
V
I
260
|
V
V
P
P
R
R
R
R
K
K
A
A
K
K
I
I
I
I
R
R
270
|
D
D
Y
Y
G
G
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
280
|
C
C
V
V
A
A
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q182H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.E11D+p.V31I+p.G70R+p.I84V+p.L101I+p.S119P+p.T122I+p.N155H+p.G163E+p.V201I+p.D256E |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 74.457 |
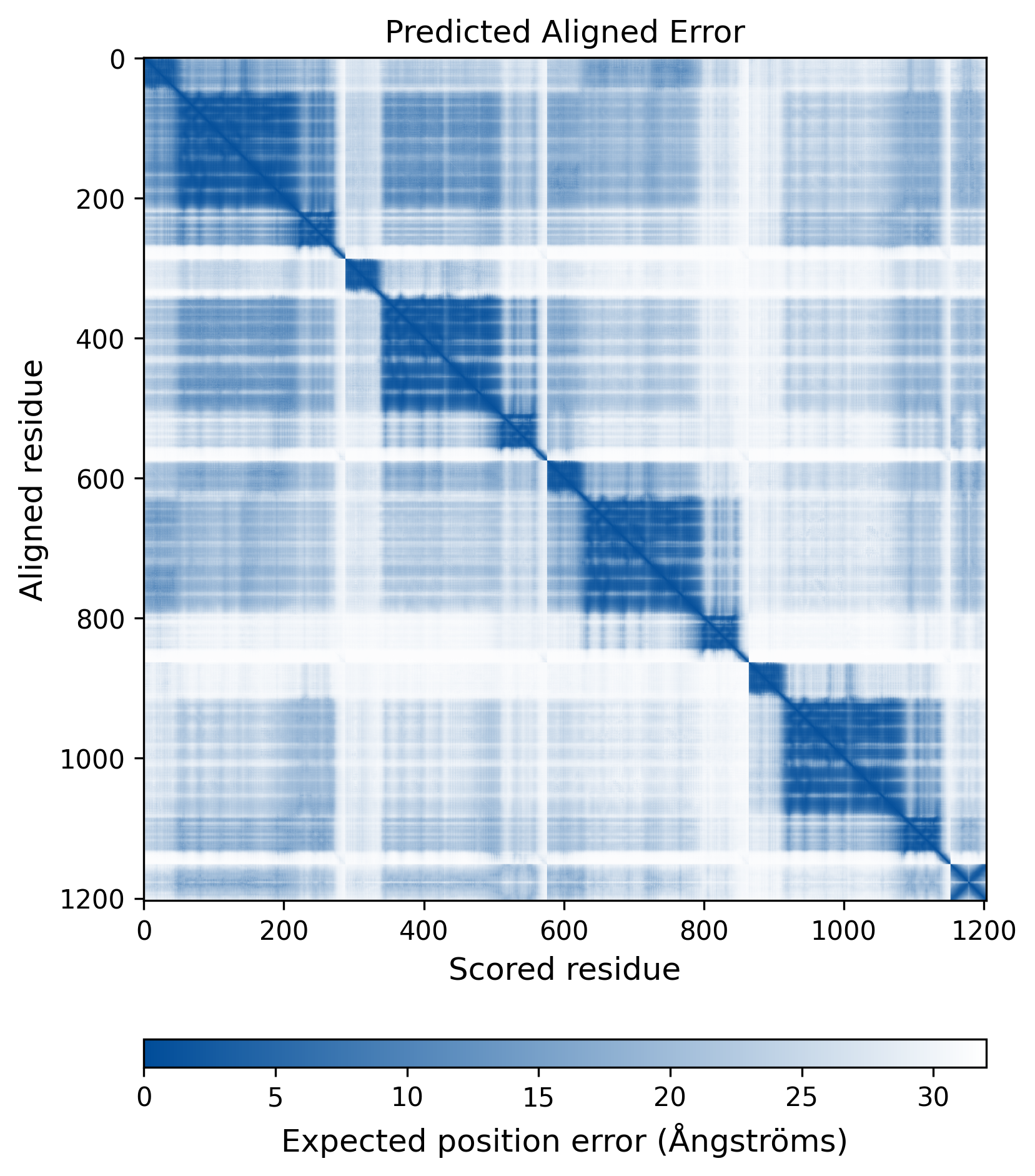
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
D
H
H
1160
|
E
E
K
K
Y
Y
H
H
S
S
N
N
W
W
R
R
A
A
M
M
1170
|
A
A
S
S
D
D
F
F
N
N
L
L
P
P
P
P
V
I
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
L
E
E
G
R
K
K
V
V
1220
|
I
I
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
V
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
F
F
L
I
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
T
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
P
N
N
F
F
T
I
1270
|
G
G
A
A
T
T
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
I
K
K
Q
Q
E
E
F
F
G
G
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
Q
G
G
V
V
V
V
E
E
1300
|
S
S
M
M
N
H
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
E
Q
Q
V
V
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
I
D
D
1350
|
I
I
I
I
A
A
T
T
D
D
I
I
Q
Q
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
S
R
R
N
N
1380
|
P
P
L
L
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
E
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
V
A
A
1430
|
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q175H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.K14R+p.S24G+p.D25E+p.S39C+p.I60IM+p.L63I+p.T97A+p.L101I+p.S119SR+p.T124N+p.T125A+p.I135V+p.E138T+p.G140S+p.Q148H+p.M154I+p.V165I+p.V201I+p.V259VI |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 74.011 |
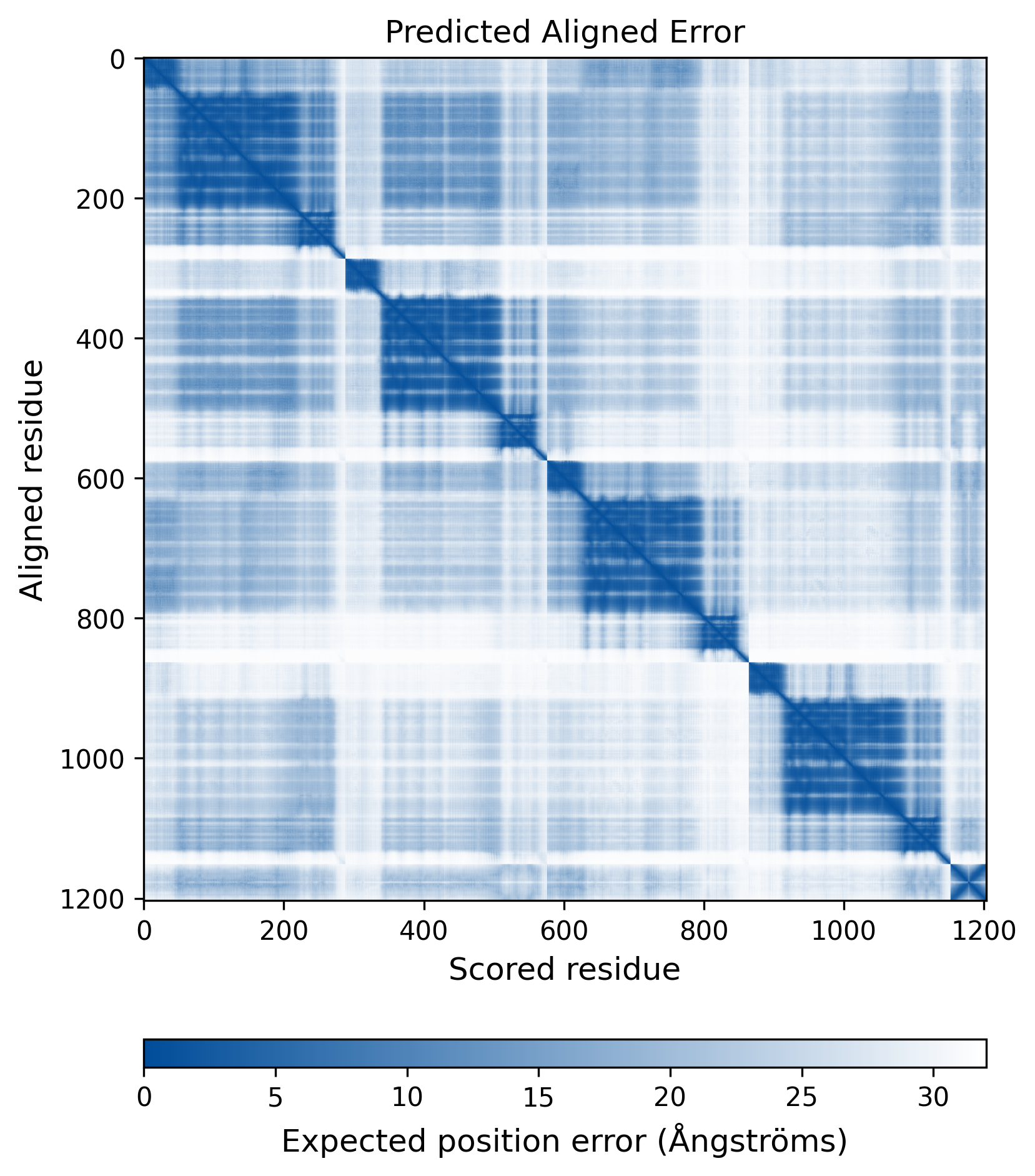
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
E
H
H
1160
|
E
E
K
R
Y
Y
H
H
S
S
N
N
W
W
R
R
A
A
M
M
1170
|
A
A
S
G
D
E
F
F
N
N
L
L
P
P
P
P
V
V
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
C
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
M
W
W
Q
Q
1210
|
L
I
D
D
C
C
T
T
H
H
L
L
E
E
G
G
K
K
V
V
1220
|
I
I
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
A
A
A
Y
Y
F
F
L
I
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
T
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
R
N
N
F
F
T
T
1270
|
G
G
T
N
T
A
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
V
K
K
Q
Q
E
T
F
F
G
S
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
H
G
G
V
V
V
V
E
E
1300
|
S
S
M
I
N
N
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
G
Q
Q
V
I
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
I
D
D
1350
|
I
I
I
I
A
A
T
T
D
D
I
I
Q
Q
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
S
R
R
N
N
1380
|
P
P
L
L
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
D
I
I
K
K
V
I
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
V
A
A
1430
|
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q183H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.E11D+p.S17C+p.L68V+p.L101I+p.T124N+p.T125A+p.I135V+p.G140A+p.Q148R+p.D253E+p.V281M |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 73.975 |
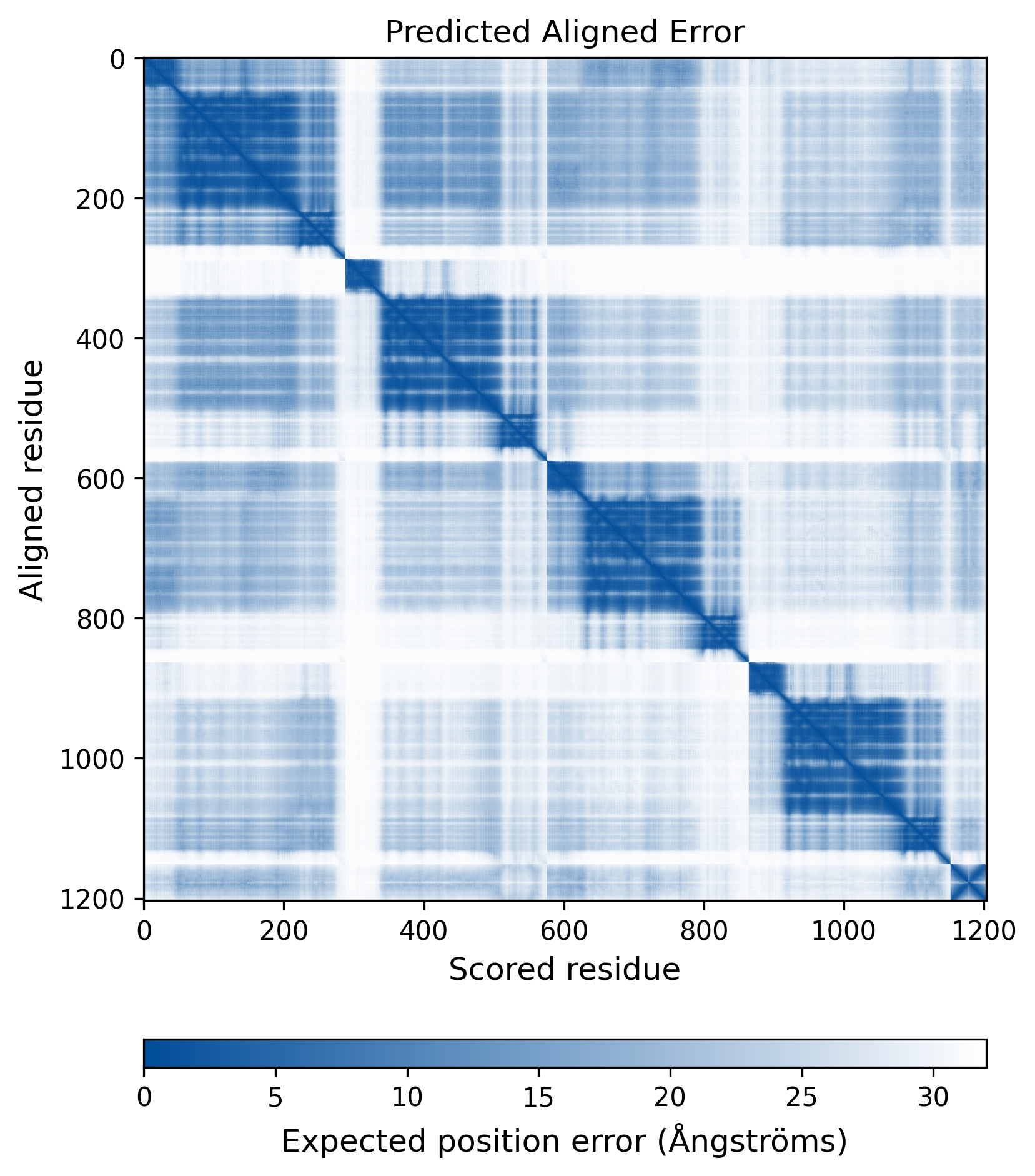
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
D
H
H
1160
|
E
E
K
K
Y
Y
H
H
S
C
N
N
W
W
R
R
A
A
M
M
1170
|
A
A
S
S
D
D
F
F
N
N
L
L
P
P
P
P
V
V
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
V
E
E
G
G
K
K
V
V
1220
|
I
I
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
F
F
L
I
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
T
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
S
N
N
F
F
T
T
1270
|
G
G
T
N
T
A
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
V
K
K
Q
Q
E
E
F
F
G
A
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
R
G
G
V
V
V
V
E
E
1300
|
S
S
M
M
N
N
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
V
D
D
1350
|
I
I
I
I
A
A
T
T
D
D
I
I
Q
Q
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
S
R
R
N
N
1380
|
P
P
L
L
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
E
N
N
S
S
D
D
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
M
A
A
1430
|
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q181H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.K14R+p.V31I+p.T97A+p.L101I+p.T112V+p.S119R+p.T124A+p.T125S+p.G134N+p.K136T+p.N155H+p.V165I+p.V201I+p.T206S+p.Y227F+p.L234I+p.S255N+p.D256E+p.S283G |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 74.006 |
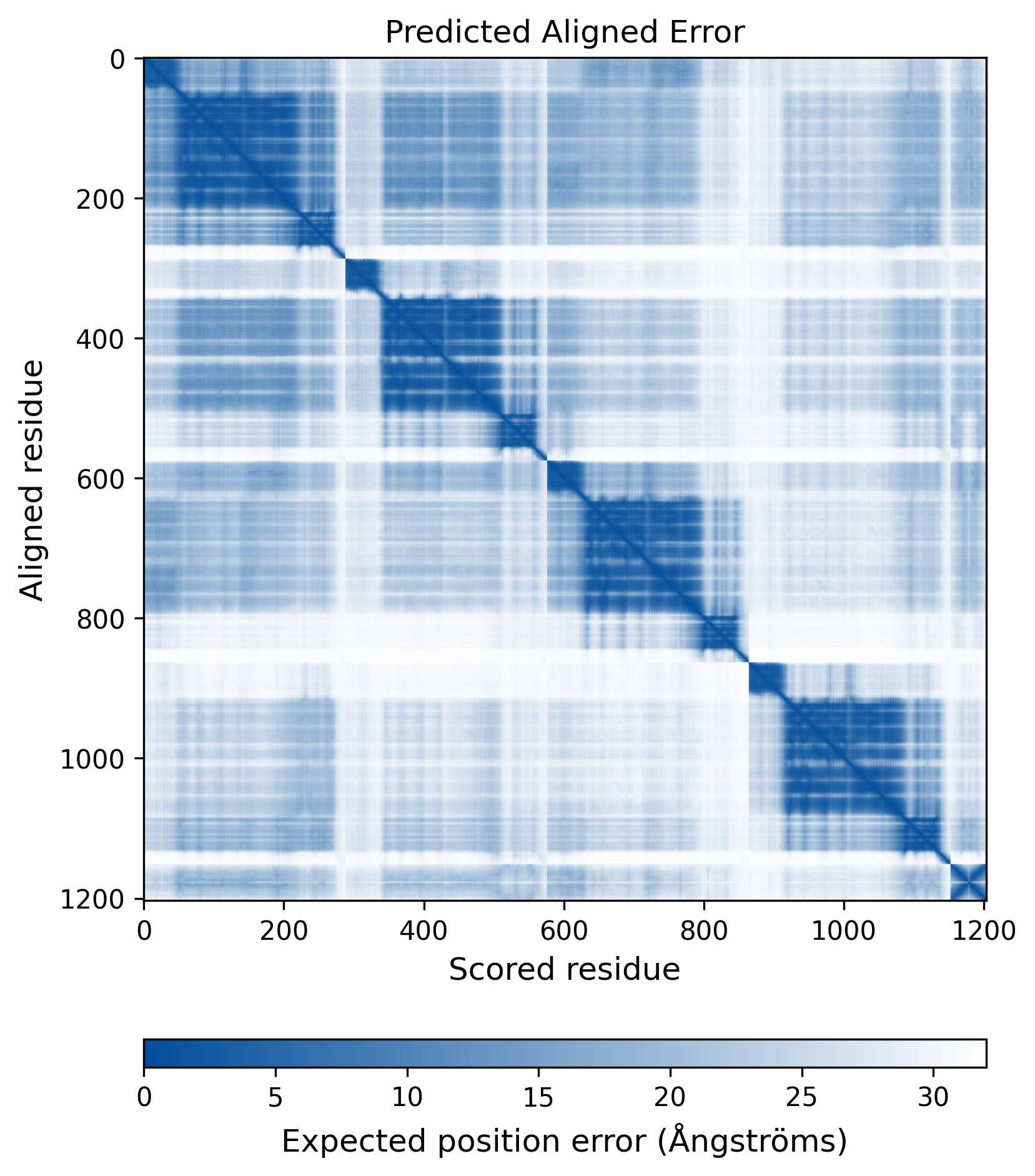
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
E
H
H
1160
|
E
E
K
R
Y
Y
H
H
S
S
N
N
W
W
R
R
A
A
M
M
1170
|
A
A
S
S
D
D
F
F
N
N
L
L
P
P
P
P
V
I
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
L
E
E
G
G
K
K
V
V
1220
|
I
I
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
A
A
A
Y
Y
F
F
L
I
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
V
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
R
N
N
F
F
T
T
1270
|
G
G
T
A
T
S
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
N
I
I
K
T
Q
Q
E
E
F
F
G
G
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
Q
G
G
V
V
V
V
E
E
1300
|
S
S
M
M
N
H
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
G
Q
Q
V
I
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
I
D
D
1350
|
I
I
I
I
A
A
T
S
D
D
I
I
Q
Q
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
F
R
R
D
D
S
S
R
R
N
N
1380
|
P
P
L
I
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
N
D
E
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
V
A
A
1430
|
S
G
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q178H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S17N+p.R20K+p.M50I+p.V77A+p.L101I+p.T112I+p.T124A+p.T125A+p.K136Q+p.G193E+p.V201I+p.T206S+p.T218L+p.I220V+p.L234V+p.D256E+p.R263K+p.D279G+p.S283G |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 75.488 |
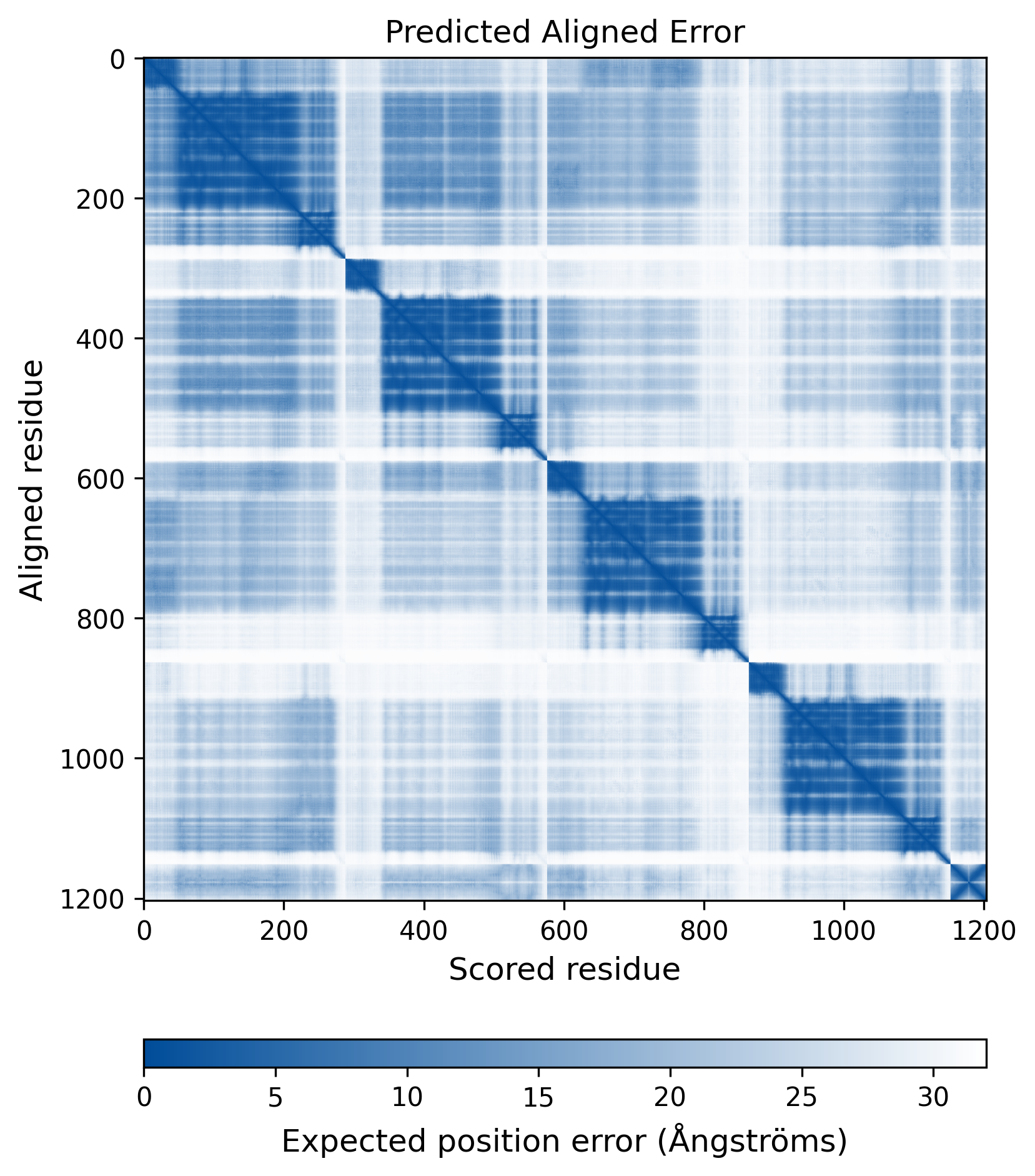
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
E
H
H
1160
|
E
E
K
K
Y
Y
H
H
S
N
N
N
W
W
R
K
A
A
M
M
1170
|
A
A
S
S
D
D
F
F
N
N
L
L
P
P
P
P
V
V
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
I
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
L
E
E
G
G
K
K
V
V
1220
|
I
I
L
L
V
V
A
A
V
A
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
F
F
L
I
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
I
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
S
N
N
F
F
T
T
1270
|
G
G
T
A
T
A
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
I
K
Q
Q
Q
E
E
F
F
G
G
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
Q
G
G
V
V
V
V
E
E
1300
|
S
S
M
M
N
N
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
E
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
I
D
D
1350
|
I
I
I
I
A
A
T
S
D
D
I
I
Q
Q
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
L
K
K
I
V
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
S
R
R
N
N
1380
|
P
P
L
V
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
E
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
K
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
G
C
C
V
V
A
A
1430
|
S
G
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q179H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.K14R+p.E69ED+p.E138K+p.G140S+p.Q148H+p.M154I+p.N155H+p.V201I |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 74.004 |
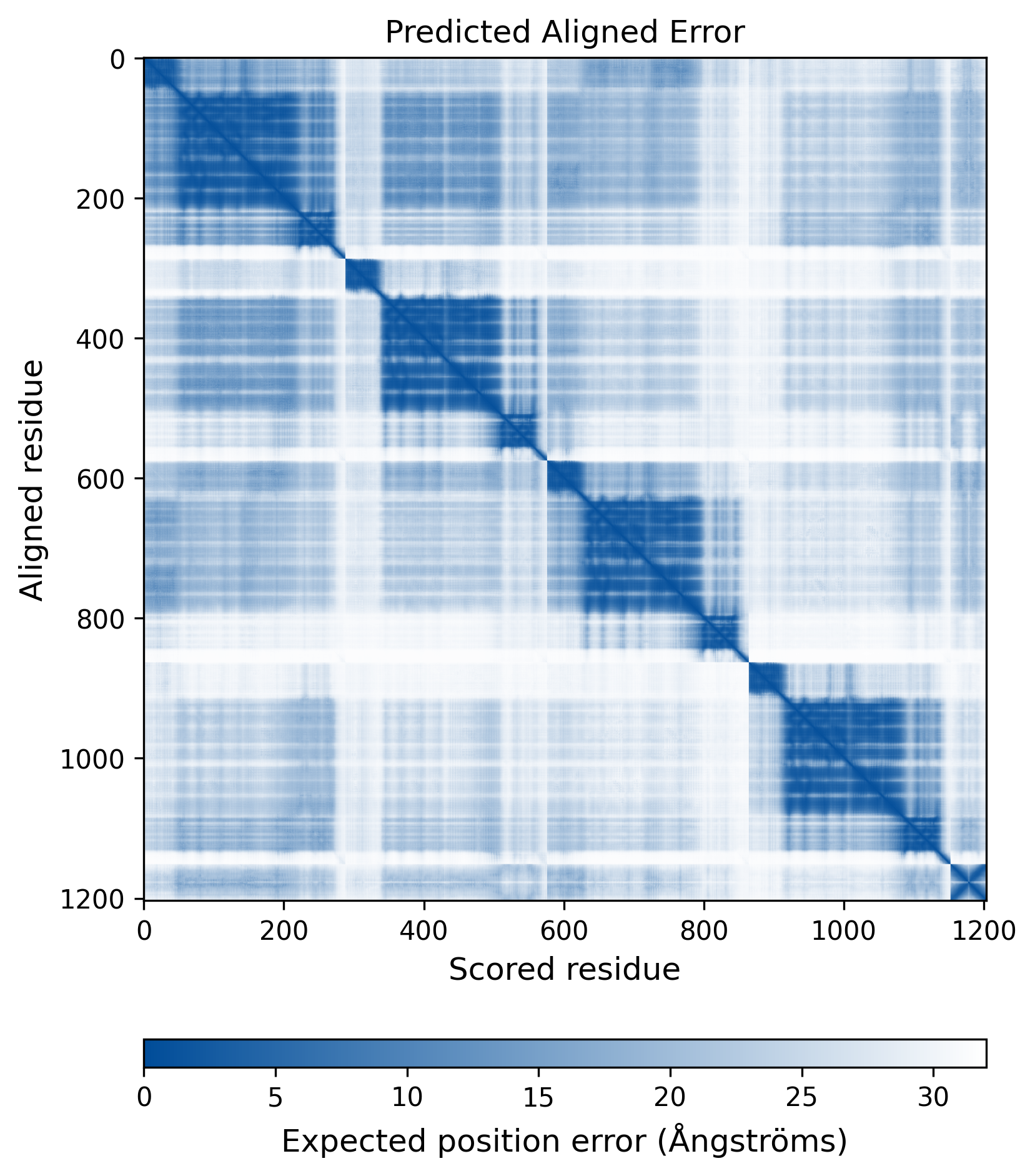
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
E
H
H
1160
|
E
E
K
R
Y
Y
H
H
S
S
N
N
W
W
R
R
A
A
M
M
1170
|
A
A
S
S
D
D
F
F
N
N
L
L
P
P
P
P
V
V
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
L
E
D
G
G
K
K
V
V
1220
|
I
I
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
F
F
L
L
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
T
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
S
N
N
F
F
T
T
1270
|
G
G
A
A
T
T
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
I
K
K
Q
Q
E
K
F
F
G
S
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
H
G
G
V
V
V
V
E
E
1300
|
S
S
M
I
N
H
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
I
D
D
1350
|
I
I
I
I
A
A
T
T
D
D
I
I
Q
Q
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
S
R
R
N
N
1380
|
P
P
L
L
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
D
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
V
A
A
1430
|
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q185H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S17N+p.D25E+p.S119G+p.T122I+p.T124N+p.T125A+p.M154I+p.N155H+p.K156R+p.T206S+p.T218S |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 74.293 |
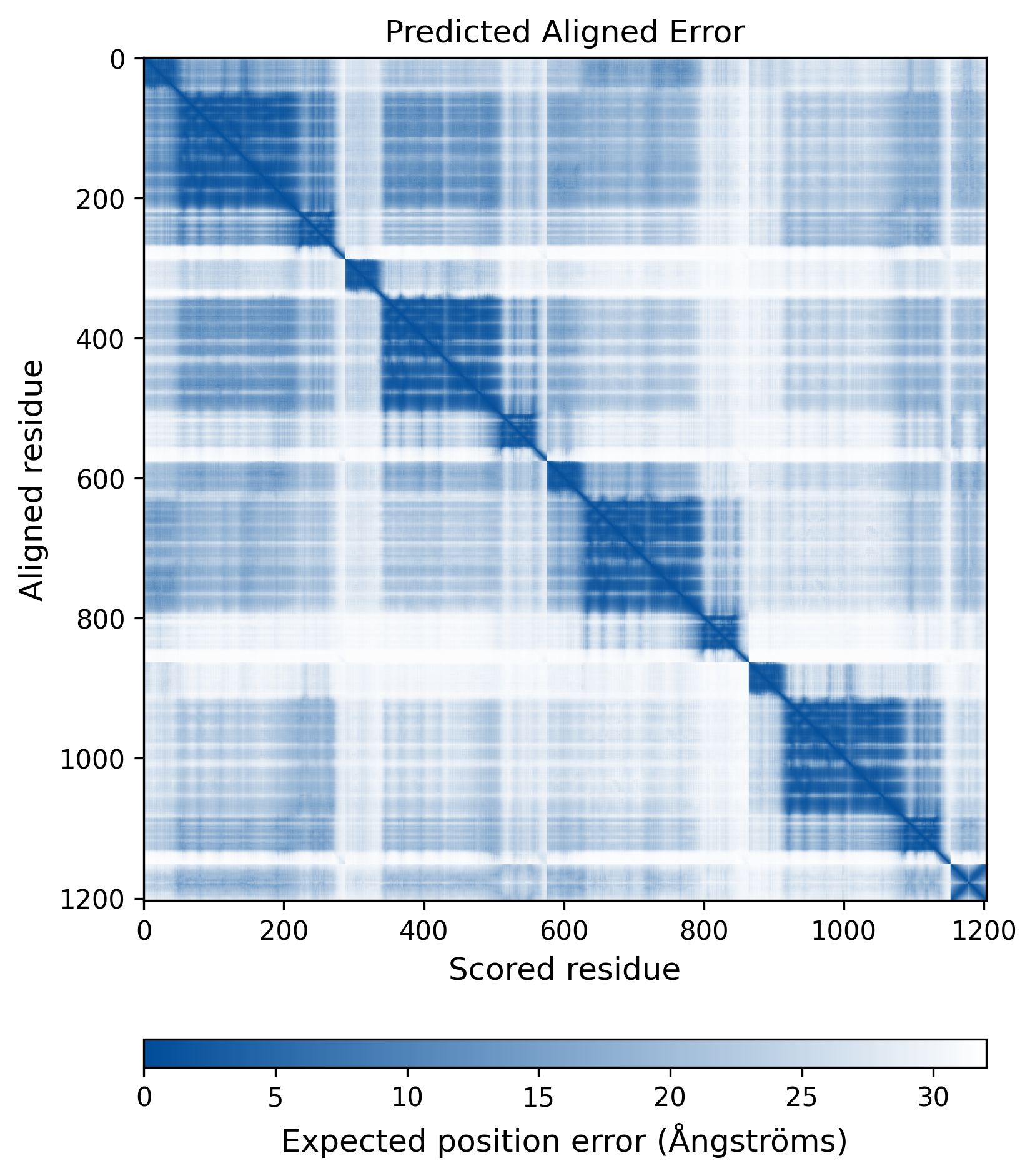
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
E
H
H
1160
|
E
E
K
K
Y
Y
H
H
S
N
N
N
W
W
R
R
A
A
M
M
1170
|
A
A
S
S
D
E
F
F
N
N
L
L
P
P
P
P
V
V
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
L
E
E
G
G
K
K
V
V
1220
|
I
I
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
F
F
L
L
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
T
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
G
N
N
F
F
T
I
1270
|
G
G
T
N
T
A
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
I
K
K
Q
Q
E
E
F
F
G
G
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
Q
G
G
V
V
V
V
E
E
1300
|
S
S
M
I
N
H
K
R
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
V
D
D
1350
|
I
I
I
I
A
A
T
S
D
D
I
I
Q
Q
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
S
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
S
R
R
N
N
1380
|
P
P
L
L
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
D
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
V
A
A
1430
|
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q177H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Resistant Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.G59D+p.I60V+p.E138K+p.Q148R+p.K173R+p.T218S+p.S230N+p.M275S+p.C280S |
|||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q180H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Drug Sensitivity Data Categorized by Their Corresponding Mechanisms | |||||||||||||
|
|
|||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Sensitive Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S17N+p.S39R+p.L45V+p.M50ML+p.T66A+p.V75I+p.L101I+p.T124A+p.K127R+p.S147SG+p.G163K+p.V201I |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 74.462 |
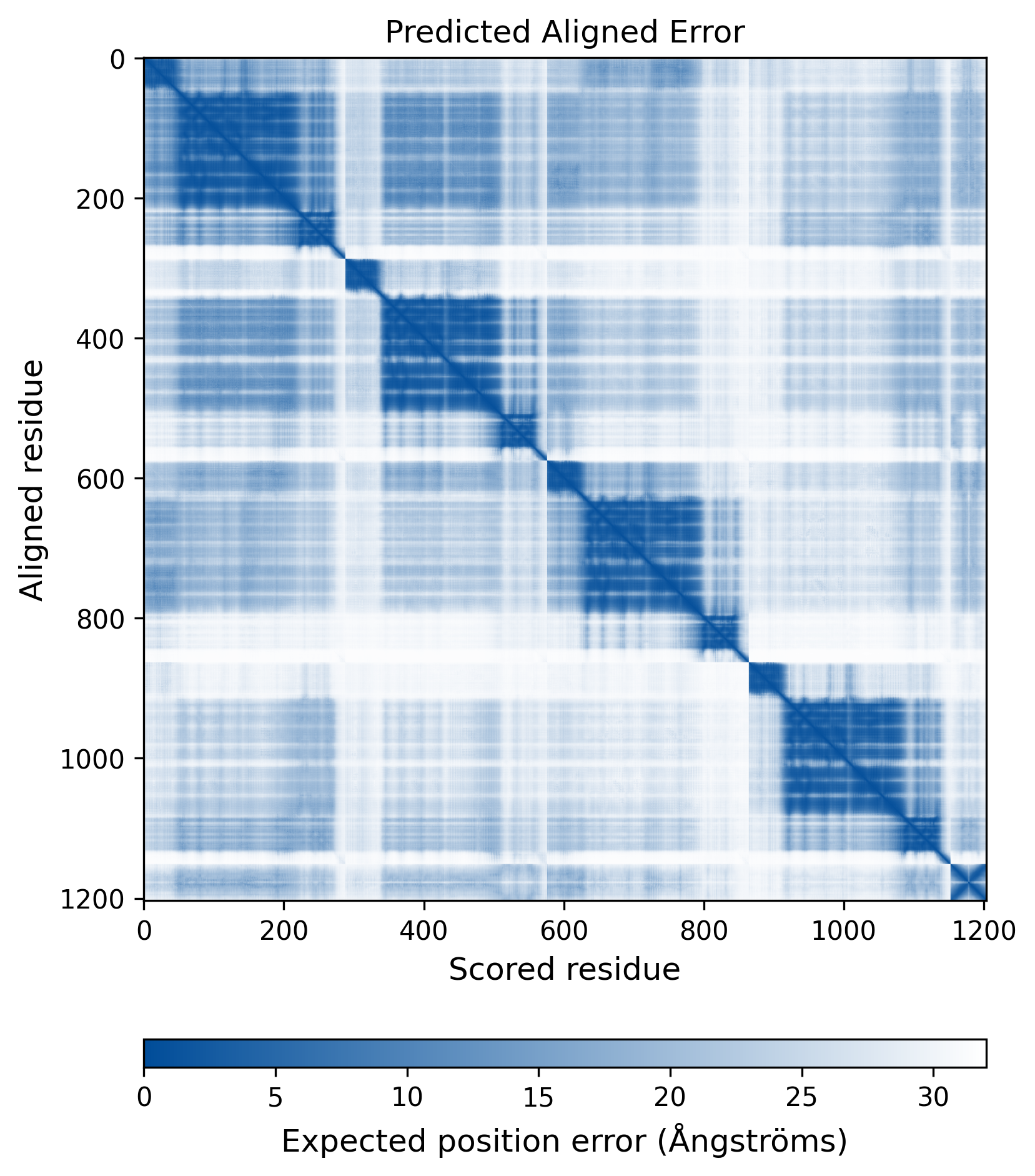
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
E
H
H
1160
|
E
E
K
K
Y
Y
H
H
S
N
N
N
W
W
R
R
A
A
M
M
1170
|
A
A
S
S
D
D
F
F
N
N
L
L
P
P
P
P
V
V
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
R
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
V
K
K
G
G
E
E
A
A
M
L
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
A
H
H
L
L
E
E
G
G
K
K
V
V
1220
|
I
I
L
L
V
I
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
F
F
L
I
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
T
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
S
N
N
F
F
T
T
1270
|
G
G
T
A
T
T
V
V
K
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
I
K
K
Q
Q
E
E
F
F
G
G
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
G
Q
Q
G
G
V
V
V
V
E
E
1300
|
S
S
M
M
N
N
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
K
Q
Q
V
V
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
I
D
D
1350
|
I
I
I
I
A
A
T
T
D
D
I
I
Q
Q
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
S
R
R
N
N
1380
|
P
P
L
L
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
D
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
V
A
A
1430
|
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q168H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Sensitive Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.A23T+p.S24G+p.D25E+p.L28I+p.I72V+p.T124A+p.Q148R+p.V165I+p.L172LF+p.H183HP+p.Q209QP+p.A239AT |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 73.352 |
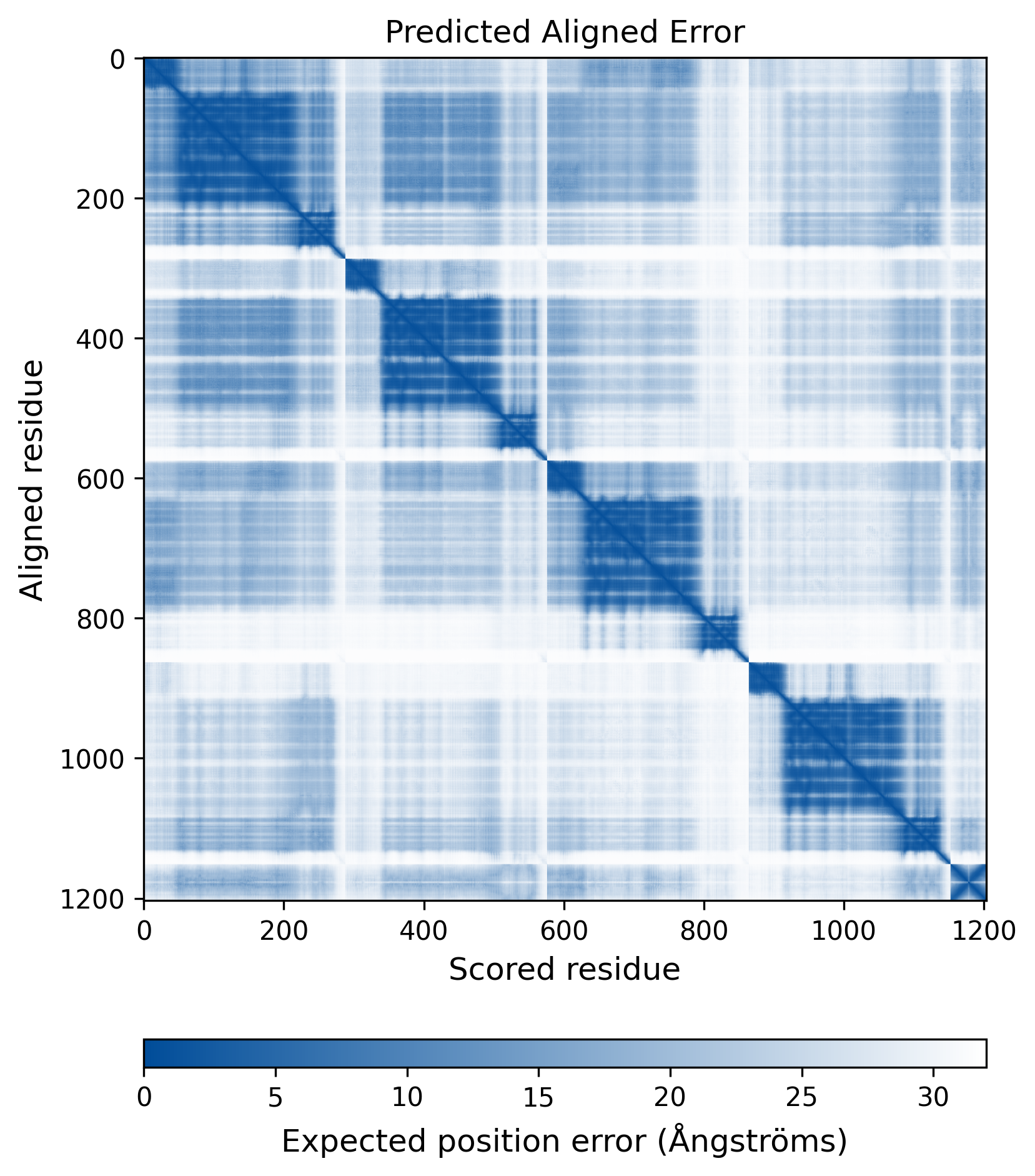
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
E
H
H
1160
|
E
E
K
K
Y
Y
H
H
S
S
N
N
W
W
R
R
A
A
M
M
1170
|
A
T
S
G
D
E
F
F
N
N
L
I
P
P
P
P
V
V
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
L
E
E
G
G
K
K
I
V
1220
|
I
I
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
F
F
L
L
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
T
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
S
N
N
F
F
T
T
1270
|
G
G
T
A
T
T
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
I
K
K
Q
Q
E
E
F
F
G
G
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
R
G
G
V
V
V
V
E
E
1300
|
S
S
M
M
N
N
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
G
Q
Q
V
I
R
R
D
D
Q
Q
A
A
E
E
H
H
L
F
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
P
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
V
D
D
1350
|
I
I
I
I
A
A
T
T
D
D
I
I
Q
P
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
S
R
R
N
N
1380
|
P
P
L
L
W
W
K
K
G
G
P
P
A
T
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
D
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
V
A
A
1430
|
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q169H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Sensitive Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.K7KR+p.S17N+p.I72V+p.I84L+p.T97A+p.L101I+p.T112I+p.S119P+p.T122I+p.T124N+p.T125A+p.K136Q+p.Y143R+p.V201I+p.A205S+p.K211R+p.T218L+p.L234V+p.D256E+p.I268L+p.D270H |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 73.893 |
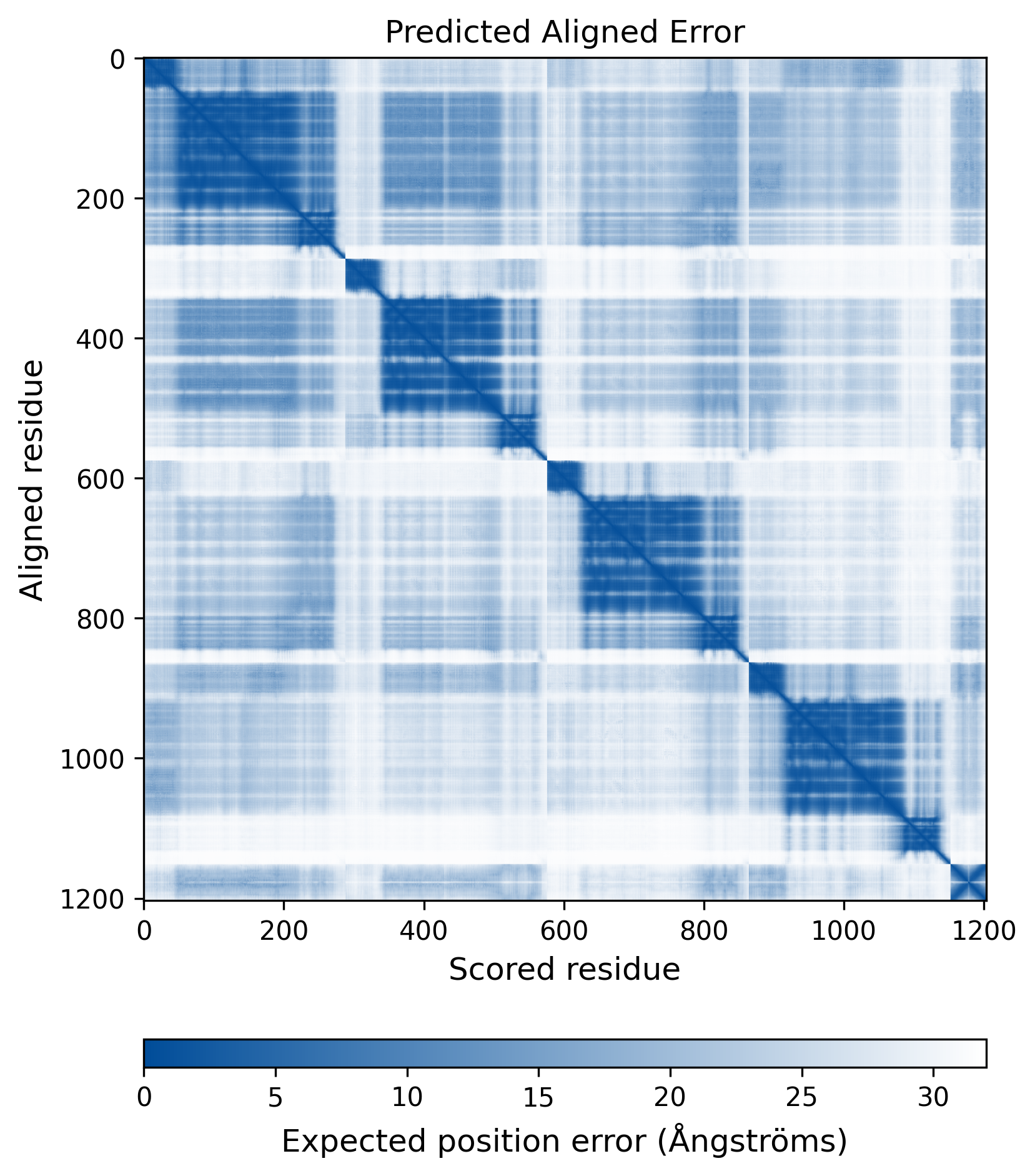
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
R
A
A
Q
Q
D
D
E
E
H
H
1160
|
E
E
K
K
Y
Y
H
H
S
N
N
N
W
W
R
R
A
A
M
M
1170
|
A
A
S
S
D
D
F
F
N
N
L
L
P
P
P
P
V
V
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
L
E
E
G
G
K
K
I
V
1220
|
I
I
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
L
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
A
A
A
Y
Y
F
F
L
I
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
I
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
P
N
N
F
F
T
I
1270
|
G
G
T
N
T
A
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
I
K
Q
Q
Q
E
E
F
F
G
G
I
I
P
P
1290
|
Y
R
N
N
P
P
Q
Q
S
S
Q
Q
G
G
V
V
V
V
E
E
1300
|
S
S
M
M
N
N
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
I
D
D
1350
|
I
I
I
I
A
S
T
T
D
D
I
I
Q
Q
T
T
K
R
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
L
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
S
R
R
N
N
1380
|
P
P
L
V
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
E
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
L
R
R
D
H
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
V
A
A
1430
|
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q170H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Sensitive Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.S17N+p.M154L+p.N155H+p.V201I+p.D232N+p.L242LF |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 73.66 |
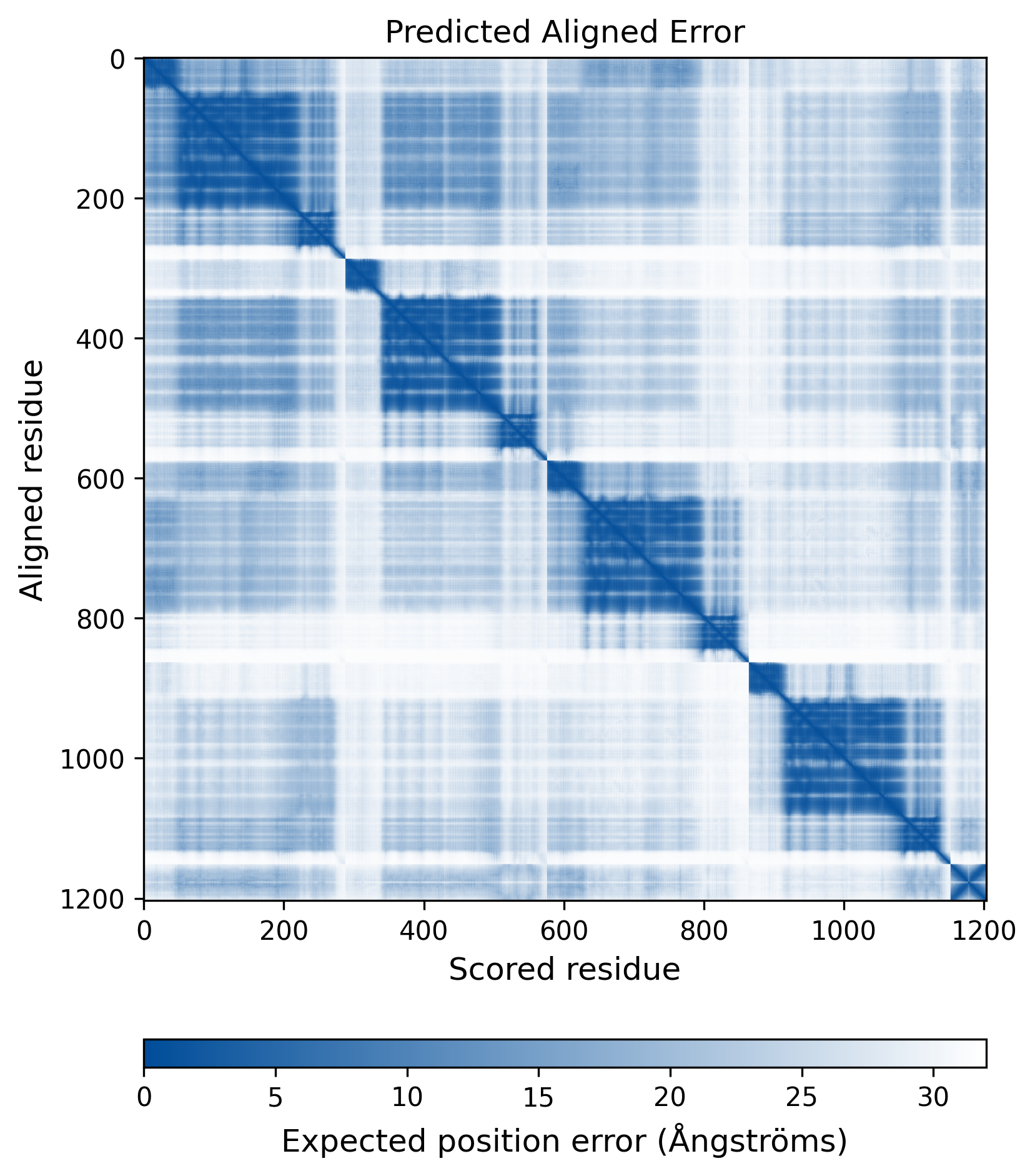
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
E
H
H
1160
|
E
E
K
K
Y
Y
H
H
S
N
N
N
W
W
R
R
A
A
M
M
1170
|
A
A
S
S
D
D
F
F
N
N
L
L
P
P
P
P
V
V
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
L
E
E
G
G
K
K
V
V
1220
|
I
I
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
F
F
L
L
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
T
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
S
N
N
F
F
T
T
1270
|
G
G
A
A
T
T
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
I
K
K
Q
Q
E
E
F
F
G
G
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
Q
G
G
V
V
V
V
E
E
1300
|
S
S
M
L
N
H
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
I
D
D
1350
|
I
I
I
I
A
A
T
T
D
D
I
I
Q
Q
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
S
R
R
D
N
1380
|
P
P
L
L
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
F
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
D
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
V
A
A
1430
|
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q171H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Sensitive Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.K14R+p.S24H+p.V31I+p.L68I+p.I73V+p.E92Q+p.L101I+p.T112V+p.S119T+p.T124G+p.T125A+p.G134N+p.K136T+p.V165I+p.V201I+p.T206S+p.L234I+p.D256E+p.S283G |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 73.995 |
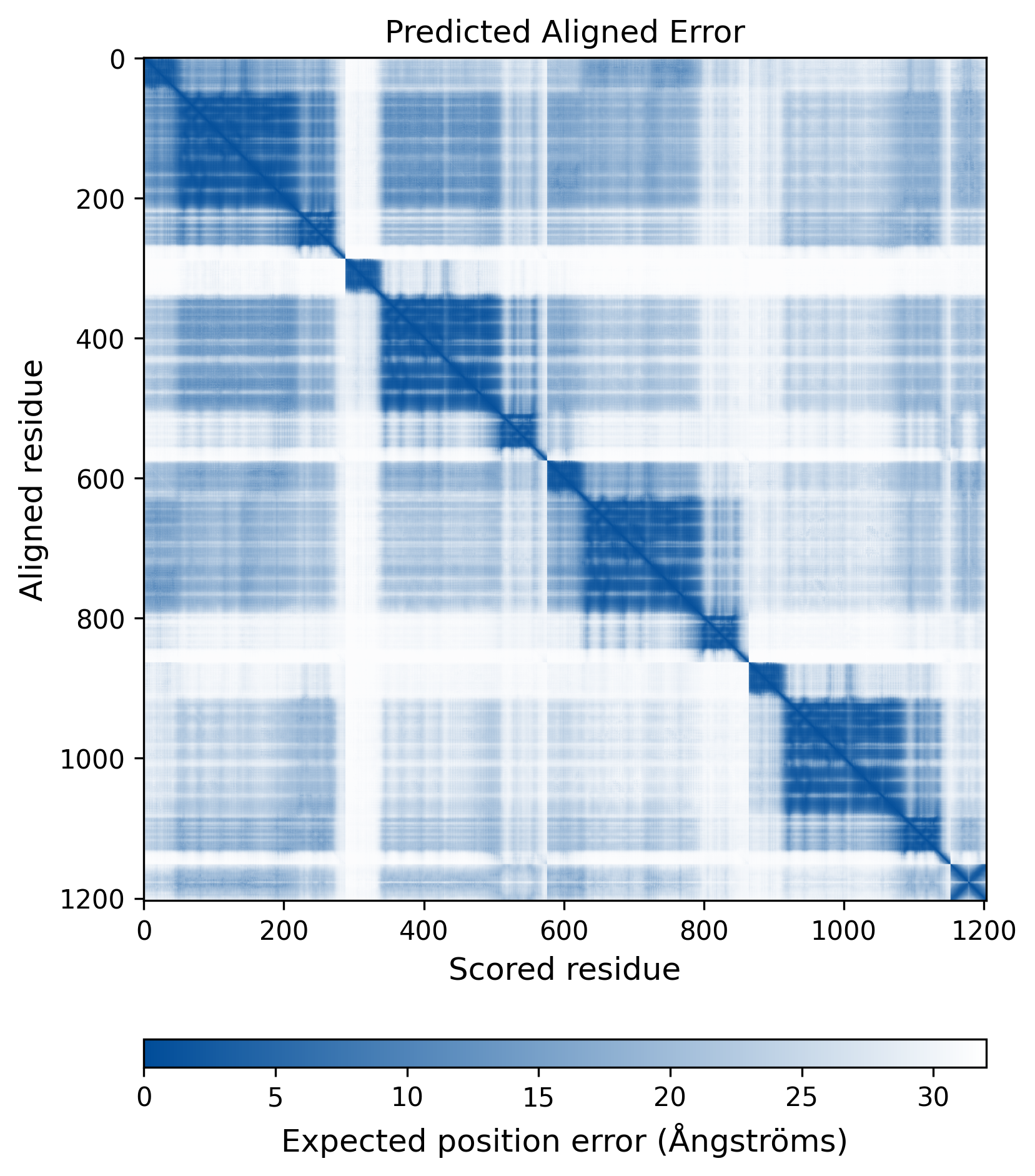
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
E
H
H
1160
|
E
E
K
R
Y
Y
H
H
S
S
N
N
W
W
R
R
A
A
M
M
1170
|
A
A
S
H
D
D
F
F
N
N
L
L
P
P
P
P
V
I
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
I
E
E
G
G
K
K
V
V
1220
|
I
V
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
Q
1240
|
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
F
F
L
I
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
V
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
T
N
N
F
F
T
T
1270
|
G
G
T
G
T
A
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
N
I
I
K
T
Q
Q
E
E
F
F
G
G
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
Q
G
G
V
V
V
V
E
E
1300
|
S
S
M
M
N
N
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
G
Q
Q
V
I
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
I
D
D
1350
|
I
I
I
I
A
A
T
S
D
D
I
I
Q
Q
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
S
R
R
N
N
1380
|
P
P
L
I
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
E
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
V
A
A
1430
|
S
G
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q167H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Sensitive Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.E11D+p.R20K+p.E92Q+p.V201I+p.K211R+p.S230SN+p.D232E+p.D256E |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 74.87 |
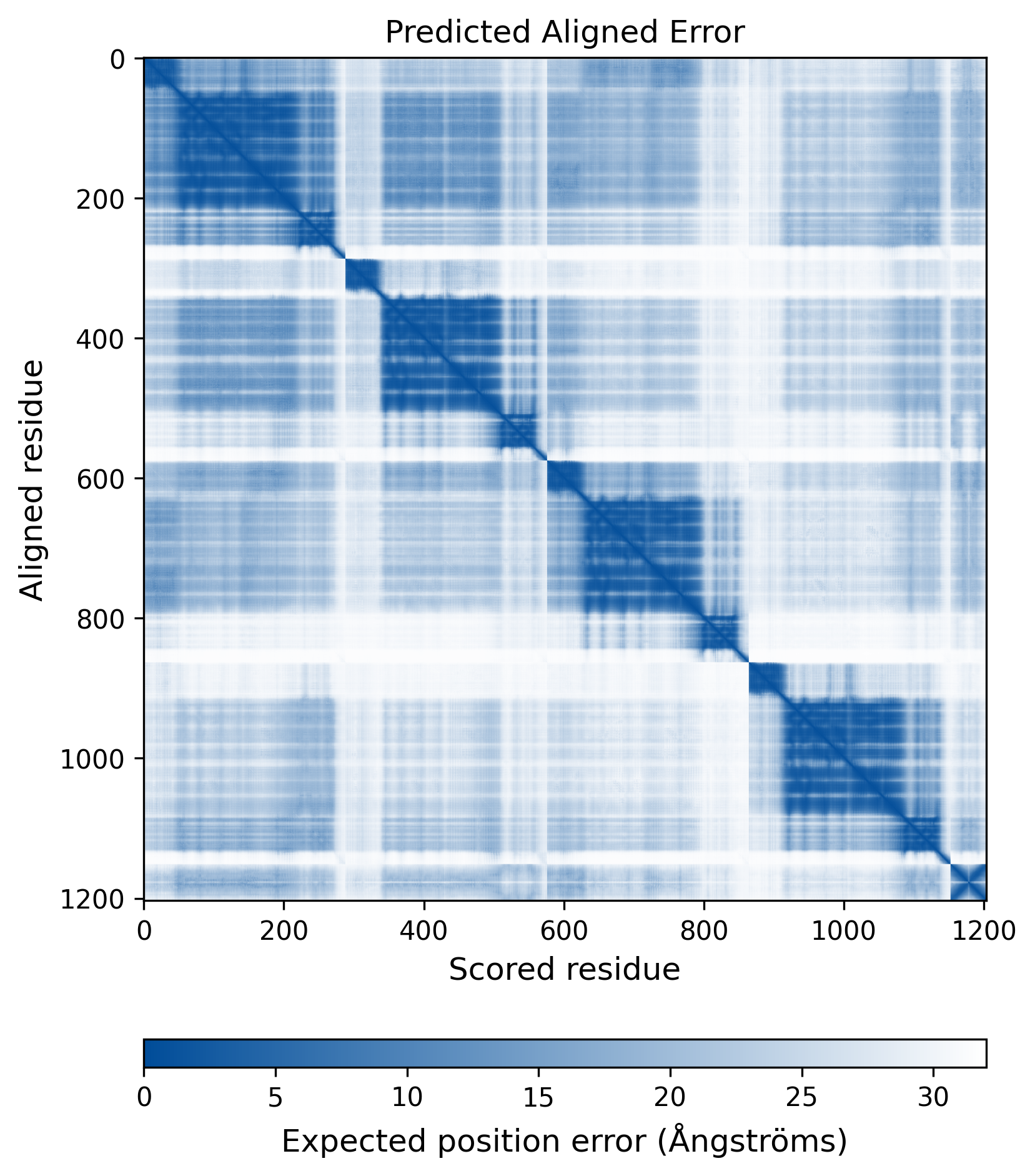
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
D
H
H
1160
|
E
E
K
K
Y
Y
H
H
S
S
N
N
W
W
R
K
A
A
M
M
1170
|
A
A
S
S
D
D
F
F
N
N
L
L
P
P
P
P
V
V
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
L
E
E
G
G
K
K
V
V
1220
|
I
I
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
Q
1240
|
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
F
F
L
L
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
T
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
S
N
N
F
F
T
T
1270
|
G
G
A
A
T
T
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
I
K
K
Q
Q
E
E
F
F
G
G
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
Q
G
G
V
V
V
V
E
E
1300
|
S
S
M
M
N
N
K
K
E
E
L
L
K
K
K
K
I
I
I
I
1310
|
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
G
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
I
D
D
1350
|
I
I
I
I
A
A
T
T
D
D
I
I
Q
Q
T
T
K
R
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
N
R
R
D
E
1380
|
P
P
L
L
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
E
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
A
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
D
C
C
V
V
A
A
1430
|
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q172H/k/R plus one to two additional INSTI mutations. | ||||||||||||
| Key Molecule: HIV1 Integrase (HIV1 IT) | [1] | ||||||||||||
| Sensitive Disease | Human immunodeficiency virus infection [ICD-11: 1C62.0] | ||||||||||||
| Molecule Alteration | Missense mutation | p.K14R+p.L101V+p.T112I+p.T125V+p.N155H+p.E157Q+p.K160Q+p.G193E+p.S230N+p.L234I+p.A265V+p.D279G |
|||||||||||
| Wild Type Structure | Method: AlphaFold | Average pLDDT: 75.736 |

|
||||||||||
| Mutant Type Structure | Method: AlphaFold | Average pLDDT: 72.839 |
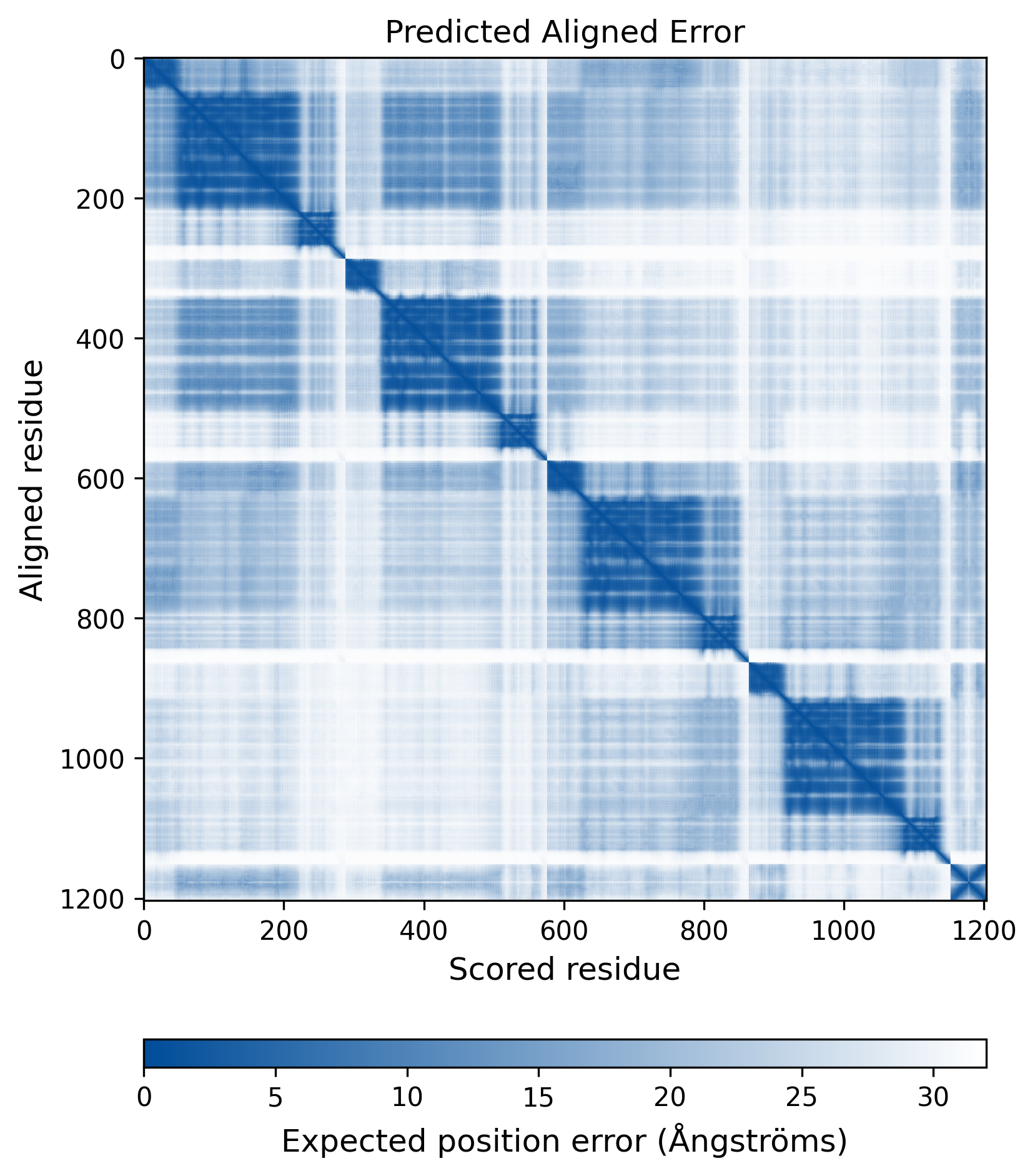
|
||||||||||
| Download The Information of Sequence | Download The Structure File | ||||||||||||
-
F
F
L
L
1150
|
D
D
G
G
I
I
D
D
K
K
A
A
Q
Q
D
D
E
E
H
H
1160
|
E
E
K
R
Y
Y
H
H
S
S
N
N
W
W
R
R
A
A
M
M
1170
|
A
A
S
S
D
D
F
F
N
N
L
L
P
P
P
P
V
V
V
V
1180
|
A
A
K
K
E
E
I
I
V
V
A
A
S
S
C
C
D
D
K
K
1190
|
C
C
Q
Q
L
L
K
K
G
G
E
E
A
A
M
M
H
H
G
G
1200
|
Q
Q
V
V
D
D
C
C
S
S
P
P
G
G
I
I
W
W
Q
Q
1210
|
L
L
D
D
C
C
T
T
H
H
L
L
E
E
G
G
K
K
V
V
1220
|
I
I
L
L
V
V
A
A
V
V
H
H
V
V
A
A
S
S
G
G
1230
|
Y
Y
I
I
E
E
A
A
E
E
V
V
I
I
P
P
A
A
E
E
1240
|
T
T
G
G
Q
Q
E
E
T
T
A
A
Y
Y
F
F
L
V
L
L
1250
|
K
K
L
L
A
A
G
G
R
R
W
W
P
P
V
V
K
K
T
I
1260
|
I
I
H
H
T
T
D
D
N
N
G
G
S
S
N
N
F
F
T
T
1270
|
G
G
A
A
T
V
V
V
R
R
A
A
A
A
C
C
W
W
W
W
1280
|
A
A
G
G
I
I
K
K
Q
Q
E
E
F
F
G
G
I
I
P
P
1290
|
Y
Y
N
N
P
P
Q
Q
S
S
Q
Q
G
G
V
V
V
V
E
E
1300
|
S
S
M
M
N
H
K
K
E
Q
L
L
K
K
K
Q
I
I
I
I
1310
|
G
G
Q
Q
V
V
R
R
D
D
Q
Q
A
A
E
E
H
H
L
L
1320
|
K
K
T
T
A
A
V
V
Q
Q
M
M
A
A
V
V
F
F
I
I
1330
|
H
H
N
N
F
F
K
K
R
R
K
K
G
G
G
G
I
I
G
G
1340
|
G
E
Y
Y
S
S
A
A
G
G
E
E
R
R
I
I
V
V
D
D
1350
|
I
I
I
I
A
A
T
T
D
D
I
I
Q
Q
T
T
K
K
E
E
1360
|
L
L
Q
Q
K
K
Q
Q
I
I
T
T
K
K
I
I
Q
Q
N
N
1370
|
F
F
R
R
V
V
Y
Y
Y
Y
R
R
D
D
S
N
R
R
N
N
1380
|
P
P
L
I
W
W
K
K
G
G
P
P
A
A
K
K
L
L
L
L
1390
|
W
W
K
K
G
G
E
E
G
G
A
A
V
V
V
V
I
I
Q
Q
1400
|
D
D
N
N
S
S
D
D
I
I
K
K
V
V
V
V
P
P
R
R
1410
|
R
R
K
K
A
V
K
K
I
I
I
I
R
R
D
D
Y
Y
G
G
1420
|
K
K
Q
Q
M
M
A
A
G
G
D
D
D
G
C
C
V
V
A
A
1430
|
S
S
R
R
Q
Q
D
D
E
E
D
D
|
|||||||||||||
| Experimental Note | Identified from the Human Clinical Data | ||||||||||||
| In Vitro Model | TZM-bl cells | Uterus | Homo sapiens (Human) | CVCL_B478 | |||||||||
| Experiment for Molecule Alteration |
Population sequencing of the integrase region assay | ||||||||||||
| Experiment for Drug Resistance |
TZM-bl cell line-based phenotypic assay | ||||||||||||
| Mechanism Description | In this study, DTG, BIC, and CAB demonstrated a comparable activity on a panel of INSTI-resistant strains isolated from patients exposed to RAL, EVG, and/or DTG, with a significantly reduced susceptibility only with the pathway Q173H/k/R plus one to two additional INSTI mutations. | ||||||||||||
References
If you find any error in data or bug in web service, please kindly report it to Dr. Sun and Dr. Yu.
