Molecule Information
General Information of the Molecule (ID: Mol00571)
| Name |
DNA-dependent catalytic protein kinase (PRKDC)
,Homo sapiens
|
||||
|---|---|---|---|---|---|
| Synonyms |
DNA-PK catalytic subunit; DNA-PKcs; DNPK1; p460; HYRC; HYRC1
Click to Show/Hide
|
||||
| Molecule Type |
Protein
|
||||
| Gene Name |
PRKDC
|
||||
| Gene ID | |||||
| Location |
chr8:47773111-47960178[-]
|
||||
| Sequence |
MAGSGAGVRCSLLRLQETLSAADRCGAALAGHQLIRGLGQECVLSSSPAVLALQTSLVFS
RDFGLLVFVRKSLNSIEFRECREEILKFLCIFLEKMGQKIAPYSVEIKNTCTSVYTKDRA AKCKIPALDLLIKLLQTFRSSRLMDEFKIGELFSKFYGELALKKKIPDTVLEKVYELLGL LGEVHPSEMINNAENLFRAFLGELKTQMTSAVREPKLPVLAGCLKGLSSLLCNFTKSMEE DPQTSREIFNFVLKAIRPQIDLKRYAVPSAGLRLFALHASQFSTCLLDNYVSLFEVLLKW CAHTNVELKKAALSALESFLKQVSNMVAKNAEMHKNKLQYFMEQFYGIIRNVDSNNKELS IAIRGYGLFAGPCKVINAKDVDFMYVELIQRCKQMFLTQTDTGDDRVYQMPSFLQSVASV LLYLDTVPEVYTPVLEHLVVMQIDSFPQYSPKMQLVCCRAIVKVFLALAAKGPVLRNCIS TVVHQGLIRICSKPVVLPKGPESESEDHRASGEVRTGKWKVPTYKDYVDLFRHLLSSDQM MDSILADEAFFSVNSSSESLNHLLYDEFVKSVLKIVEKLDLTLEIQTVGEQENGDEAPGV WMIPTSDPAANLHPAKPKDFSAFINLVEFCREILPEKQAEFFEPWVYSFSYELILQSTRL PLISGFYKLLSITVRNAKKIKYFEGVSPKSLKHSPEDPEKYSCFALFVKFGKEVAVKMKQ YKDELLASCLTFLLSLPHNIIELDVRAYVPALQMAFKLGLSYTPLAEVGLNALEEWSIYI DRHVMQPYYKDILPCLDGYLKTSALSDETKNNWEVSALSRAAQKGFNKVVLKHLKKTKNL SSNEAISLEEIRIRVVQMLGSLGGQINKNLLTVTSSDEMMKSYVAWDREKRLSFAVPFRE MKPVIFLDVFLPRVTELALTASDRQTKVAACELLHSMVMFMLGKATQMPEGGQGAPPMYQ LYKRTFPVLLRLACDVDQVTRQLYEPLVMQLIHWFTNNKKFESQDTVALLEAILDGIVDP VDSTLRDFCGRCIREFLKWSIKQITPQQQEKSPVNTKSLFKRLYSLALHPNAFKRLGASL AFNNIYREFREEESLVEQFVFEALVIYMESLALAHADEKSLGTIQQCCDAIDHLCRIIEK KHVSLNKAKKRRLPRGFPPSASLCLLDLVKWLLAHCGRPQTECRHKSIELFYKFVPLLPG NRSPNLWLKDVLKEEGVSFLINTFEGGGCGQPSGILAQPTLLYLRGPFSLQATLCWLDLL LAALECYNTFIGERTVGALQVLGTEAQSSLLKAVAFFLESIAMHDIIAAEKCFGTGAAGN RTSPQEGERYNYSKCTVVVRIMEFTTTLLNTSPEGWKLLKKDLCNTHLMRVLVQTLCEPA SIGFNIGDVQVMAHLPDVCVNLMKALKMSPYKDILETHLREKITAQSIEELCAVNLYGPD AQVDRSRLAAVVSACKQLHRAGLLHNILPSQSTDLHHSVGTELLSLVYKGIAPGDERQCL PSLDLSCKQLASGLLELAFAFGGLCERLVSLLLNPAVLSTASLGSSQGSVIHFSHGEYFY SLFSETINTELLKNLDLAVLELMQSSVDNTKMVSAVLNGMLDQSFRERANQKHQGLKLAT TILQHWKKCDSWWAKDSPLETKMAVLALLAKILQIDSSVSFNTSHGSFPEVFTTYISLLA DTKLDLHLKGQAVTLLPFFTSLTGGSLEELRRVLEQLIVAHFPMQSREFPPGTPRFNNYV DCMKKFLDALELSQSPMLLELMTEVLCREQQHVMEELFQSSFRRIARRGSCVTQVGLLES VYEMFRKDDPRLSFTRQSFVDRSLLTLLWHCSLDALREFFSTIVVDAIDVLKSRFTKLNE STFDTQITKKMGYYKILDVMYSRLPKDDVHAKESKINQVFHGSCITEGNELTKTLIKLCY DAFTENMAGENQLLERRRLYHCAAYNCAISVICCVFNELKFYQGFLFSEKPEKNLLIFEN LIDLKRRYNFPVEVEVPMERKKKYIEIRKEAREAANGDSDGPSYMSSLSYLADSTLSEEM SQFDFSTGVQSYSYSSQDPRPATGRFRRREQRDPTVHDDVLELEMDELNRHECMAPLTAL VKHMHRSLGPPQGEEDSVPRDLPSWMKFLHGKLGNPIVPLNIRLFLAKLVINTEEVFRPY AKHWLSPLLQLAASENNGGEGIHYMVVEIVATILSWTGLATPTGVPKDEVLANRLLNFLM KHVFHPKRAVFRHNLEIIKTLVECWKDCLSIPYRLIFEKFSGKDPNSKDNSVGIQLLGIV MANDLPPYDPQCGIQSSEYFQALVNNMSFVRYKEVYAAAAEVLGLILRYVMERKNILEES LCELVAKQLKQHQNTMEDKFIVCLNKVTKSFPPLADRFMNAVFFLLPKFHGVLKTLCLEV VLCRVEGMTELYFQLKSKDFVQVMRHRDDERQKVCLDIIYKMMPKLKPVELRELLNPVVE FVSHPSTTCREQMYNILMWIHDNYRDPESETDNDSQEIFKLAKDVLIQGLIDENPGLQLI IRNFWSHETRLPSNTLDRLLALNSLYSPKIEVHFLSLATNFLLEMTSMSPDYPNPMFEHP LSECEFQEYTIDSDWRFRSTVLTPMFVETQASQGTLQTRTQEGSLSARWPVAGQIRATQQ QHDFTLTQTADGRSSFDWLTGSSTDPLVDHTSPSSDSLLFAHKRSERLQRAPLKSVGPDF GKKRLGLPGDEVDNKVKGAAGRTDLLRLRRRFMRDQEKLSLMYARKGVAEQKREKEIKSE LKMKQDAQVVLYRSYRHGDLPDIQIKHSSLITPLQAVAQRDPIIAKQLFSSLFSGILKEM DKFKTLSEKNNITQKLLQDFNRFLNTTFSFFPPFVSCIQDISCQHAALLSLDPAAVSAGC LASLQQPVGIRLLEEALLRLLPAELPAKRVRGKARLPPDVLRWVELAKLYRSIGEYDVLR GIFTSEIGTKQITQSALLAEARSDYSEAAKQYDEALNKQDWVDGEPTEAEKDFWELASLD CYNHLAEWKSLEYCSTASIDSENPPDLNKIWSEPFYQETYLPYMIRSKLKLLLQGEADQS LLTFIDKAMHGELQKAILELHYSQELSLLYLLQDDVDRAKYYIQNGIQSFMQNYSSIDVL LHQSRLTKLQSVQALTEIQEFISFISKQGNLSSQVPLKRLLNTWTNRYPDAKMDPMNIWD DIITNRCFFLSKIEEKLTPLPEDNSMNVDQDGDPSDRMEVQEQEEDISSLIRSCKFSMKM KMIDSARKQNNFSLAMKLLKELHKESKTRDDWLVSWVQSYCRLSHCRSRSQGCSEQVLTV LKTVSLLDENNVSSYLSKNILAFRDQNILLGTTYRIIANALSSEPACLAEIEEDKARRIL ELSGSSSEDSEKVIAGLYQRAFQHLSEAVQAAEEEAQPPSWSCGPAAGVIDAYMTLADFC DQQLRKEEENASVIDSAELQAYPALVVEKMLKALKLNSNEARLKFPRLLQIIERYPEETL SLMTKEISSVPCWQFISWISHMVALLDKDQAVAVQHSVEEITDNYPQAIVYPFIISSESY SFKDTSTGHKNKEFVARIKSKLDQGGVIQDFINALDQLSNPELLFKDWSNDVRAELAKTP VNKKNIEKMYERMYAALGDPKAPGLGAFRRKFIQTFGKEFDKHFGKGGSKLLRMKLSDFN DITNMLLLKMNKDSKPPGNLKECSPWMSDFKVEFLRNELEIPGQYDGRGKPLPEYHVRIA GFDERVTVMASLRRPKRIIIRGHDEREHPFLVKGGEDLRQDQRVEQLFQVMNGILAQDSA CSQRALQLRTYSVVPMTSRLGLIEWLENTVTLKDLLLNTMSQEEKAAYLSDPRAPPCEYK DWLTKMSGKHDVGAYMLMYKGANRTETVTSFRKRESKVPADLLKRAFVRMSTSPEAFLAL RSHFASSHALICISHWILGIGDRHLNNFMVAMETGGVIGIDFGHAFGSATQFLPVPELMP FRLTRQFINLMLPMKETGLMYSIMVHALRAFRSDPGLLTNTMDVFVKEPSFDWKNFEQKM LKKGGSWIQEINVAEKNWYPRQKICYAKRKLAGANPAVITCDELLLGHEKAPAFRDYVAV ARGSKDHNIRAQEPESGLSEETQVKCLMDQATDPNILGRTWEGWEPWM Click to Show/Hide
|
||||
| 3D-structure |
|
||||
| Function |
Serine/threonine-protein kinase that acts as a molecular sensor for DNA damage. Involved in DNA non-homologous end joining (NHEJ) required for double-strand break (DSB) repair and V(D)J recombination. Must be bound to DNA to express its catalytic properties. Promotes processing of hairpin DNA structures in V(D)J recombination by activation of the hairpin endonuclease artemis (DCLRE1C). Recruited by XRCC5 and XRCC6 to DNA ends and is required to protect and align broken ends of DNA, thereby preventing their degradation, and sequester the DSB for repair by NHEJ. Act as a scaffold protein to aid the localization of DNA repair proteins to the site of damage. The assembly of the DNA-PK complex at DNA ends is also required for the NHEJ ligation step. Found at the ends of chromosomes, suggesting a further role in the maintenance of telomeric stability and the prevention of chromosomal end fusion. Also involved in modulation of transcription. As part of the DNA-PK complex, involved in the early steps of ribosome assembly by promoting the processing of precursor rRNA into mature 18S rRNA in the small-subunit processome. Binding to U3 small nucleolar RNA, recruits PRKDC and XRCC5/Ku86 to the small-subunit processome. Recognizes the substrate consensus sequence [ST]-Q. Phosphorylates 'Ser-139' of histone variant H2AX, thereby regulating DNA damage response mechanism. Phosphorylates DCLRE1C, c-Abl/ABL1, histone H1, HSPCA, c-jun/JUN, p53/TP53, PARP1, POU2F1, DHX9, FH, SRF, NHEJ1/XLF, XRCC1, XRCC4, XRCC5, XRCC6, WRN, MYC and RFA2. Can phosphorylate C1D not only in the presence of linear DNA but also in the presence of supercoiled DNA. Ability to phosphorylate p53/TP53 in the presence of supercoiled DNA is dependent on C1D. Contributes to the determination of the circadian period length by antagonizing phosphorylation of CRY1 'Ser-588' and increasing CRY1 protein stability, most likely through an indirect mechanism. Plays a role in the regulation of DNA virus-mediated innate immune response by assembling into the HDP-RNP complex, a complex that serves as a platform for IRF3 phosphorylation and subsequent innate immune response activation through the cGAS-STING pathway. Also regulates the cGAS-STING pathway by catalyzing phosphorylation of CGAS, thereby impairing CGAS oligomerization and activation.
Click to Show/Hide
|
||||
| Uniprot ID | |||||
| Ensembl ID | |||||
| HGNC ID | |||||
| Click to Show/Hide the Complete Species Lineage | |||||
Type(s) of Resistant Mechanism of This Molecule
Drug Resistance Data Categorized by Drug
Approved Drug(s)
2 drug(s) in total
| Drug Sensitivity Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Disease Class: Melanoma [ICD-11: 2C30.0] | [1] | |||
| Sensitive Disease | Melanoma [ICD-11: 2C30.0] | |||
| Sensitive Drug | Cisplatin | |||
| Molecule Alteration | Expression | Down-regulation |
||
| Experimental Note | Revealed Based on the Cell Line Data | |||
| In Vitro Model | A375 cells | Skin | Homo sapiens (Human) | CVCL_0132 |
| Sk-Mel28 cells | Skin | Homo sapiens (Human) | CVCL_0526 | |
| B16 cells | Skin | Homo sapiens (Human) | CVCL_F936 | |
| HEMn-LP cells | Skin | Homo sapiens (Human) | N.A. | |
| WM451 cells | Skin | Homo sapiens (Human) | CVCL_6357 | |
| In Vivo Model | Nude mouse xenograft model | Mus musculus | ||
| Experiment for Molecule Alteration |
qRT-PCR; Western blot analysis | |||
| Experiment for Drug Resistance |
CCK8 assay; Flow cytometric analysis | |||
| Mechanism Description | microRNA-488-3p sensitizes malignant melanoma cells to cisplatin by targeting PRkDC. | |||
| Drug Sensitivity Data Categorized by Their Corresponding Mechanisms | ||||
|
|
||||
| Disease Class: Pancreatic cancer [ICD-11: 2C10.3] | [2] | |||
| Sensitive Disease | Pancreatic cancer [ICD-11: 2C10.3] | |||
| Sensitive Drug | Gemcitabine | |||
| Molecule Alteration | Expression | Down-regulation |
||
| Experimental Note | Revealed Based on the Cell Line Data | |||
| In Vitro Model | PANC-1 cells | Pancreas | Homo sapiens (Human) | CVCL_0480 |
| Experiment for Molecule Alteration |
Western blot analysis | |||
| Experiment for Drug Resistance |
MTT assay; Annexin V apoptosis assay; Caspase-3 activity assay | |||
| Mechanism Description | microRNA-101 silences RNA-Pkcs and sensitizes pancreatic cancer cells to gemcitabine. AntagomiR101 expression causes RNA-Pkcs upregulation and gemcitabine resistance. miR101 expression inhibits Akt activation in PANC-1 cells. | |||
Disease- and Tissue-specific Abundances of This Molecule
ICD Disease Classification 02

| Differential expression of molecule in resistant diseases | ||
| The Studied Tissue | Pancreas | |
| The Specified Disease | Pancreatic cancer | |
| The Expression Level of Disease Section Compare with the Healthy Individual Tissue | p-value: 5.60E-02; Fold-change: 4.40E-01; Z-score: 2.65E-01 | |
| The Expression Level of Disease Section Compare with the Adjacent Tissue | p-value: 1.56E-02; Fold-change: 5.67E-01; Z-score: 9.25E-01 | |
|
Molecule expression in the normal tissue adjacent to the diseased tissue of patients
Molecule expression in the diseased tissue of patients
Molecule expression in the normal tissue of healthy individuals
|
||
| Disease-specific Molecule Abundances |

|
Click to View the Clearer Original Diagram |
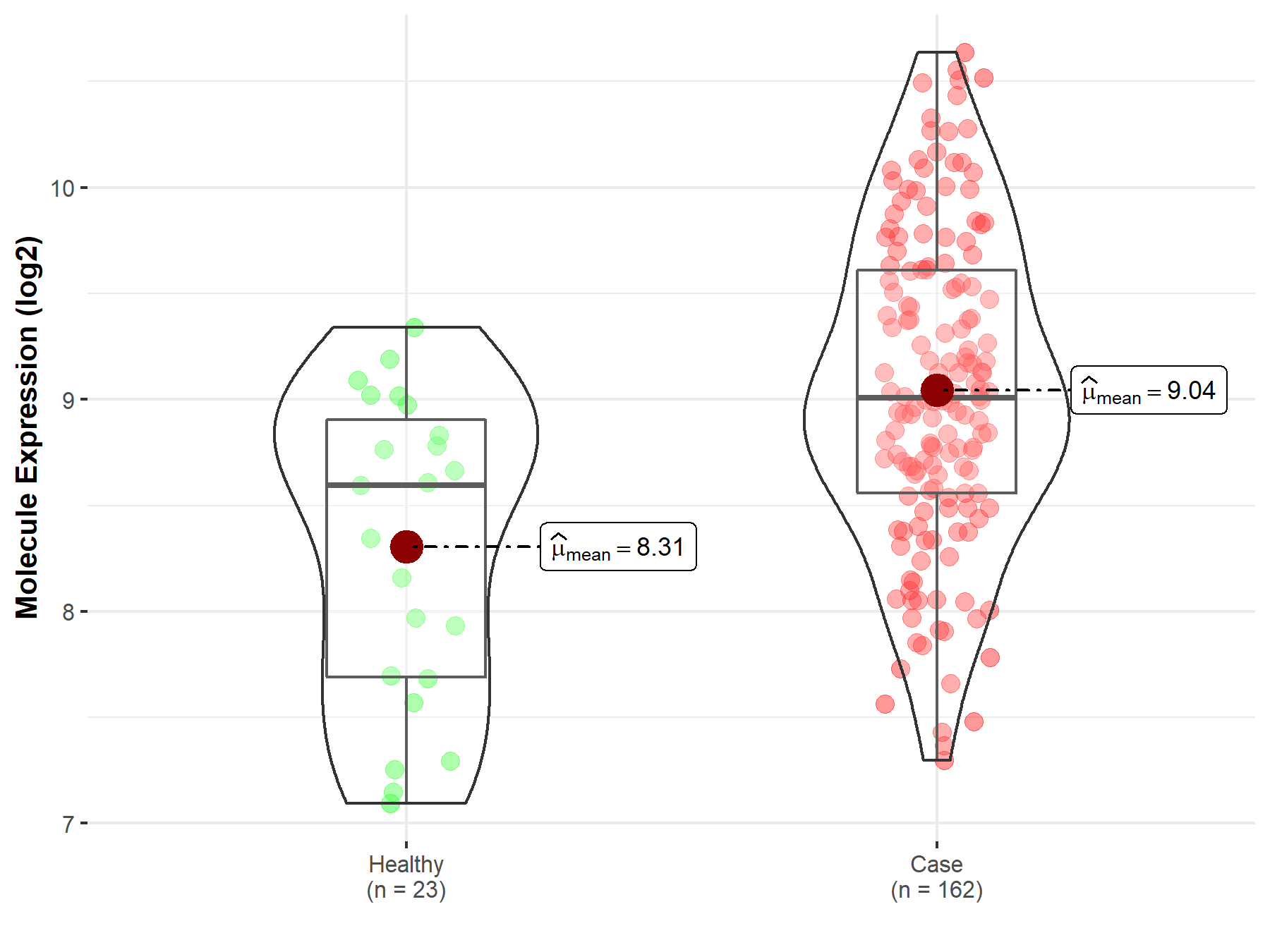
| Differential expression of molecule in resistant diseases | ||
| The Studied Tissue | Skin | |
| The Specified Disease | Melanoma | |
| The Expression Level of Disease Section Compare with the Healthy Individual Tissue | p-value: 8.27E-05; Fold-change: 4.10E-01; Z-score: 5.68E-01 | |
|
Molecule expression in the diseased tissue of patients
Molecule expression in the normal tissue of healthy individuals
|
||
| Disease-specific Molecule Abundances |
|
Click to View the Clearer Original Diagram |
Tissue-specific Molecule Abundances in Healthy Individuals

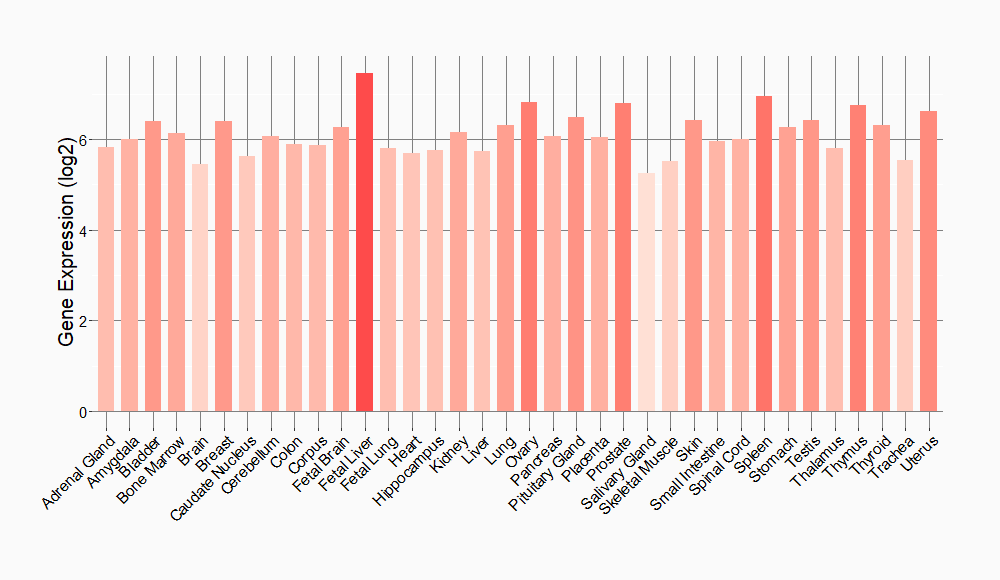
|
||
References
If you find any error in data or bug in web service, please kindly report it to Dr. Sun and Dr. Yu.
